UKCA Chemistry and Aerosol vn10.4 Tutorial 11
UKCA Chemistry and Aerosol Tutorials at vn10.4
What you will learn in this Tutorial
Before you start this tutorial, you should cd into your $HOME/roses/[SUITE-ID] directory and fcm commit your changes.
You should complete this Tutorial using a copy of your working (& committed suite) from Tutorial 10, so please copy this suite now and work from your new copy.
In this tutorial you will learn about the GLOMAP-mode aerosol module and how it tracks different aerosol types within several size classes. You will understand the standard configuration used in the UKCA jobs so far whereby the mass mixing ratios of sulphate, sea-salt, black carbon and organic matter in each mode are transported via separate tracers. GLOMAP-mode is an aerosol microphysics scheme and therefore, as well as transporting the mass of several components in the modes, the scheme also transports the number concentrations of particles in each mode.
Task 10 already introduced the basic concepts behind the GLOMAP-mode aerosol microphysics scheme and how it differs from the mass-based CLASSIC scheme which preceded UKCA.
Initially developed in the TOMCAT CTM environment (see Manktelow et al., 2007; Mann et al., 2010; Mann et al., 2012), the GLOMAP code then became the aerosol module for the UKCA sub-model of the UM (see Bellouin et al., 2013; Kipling et al., 2013; West et al., 2014; Mann et al., 2014; Dhomse et al., 2014; Turnock et al., 2015; Turnock et al., 2016; Zanchettin et al., 2016).
GLOMAP is now also implemented into the ECMWF Integrated Forecasting System as part of the "Composition IFS" module (C-IFS) where it will be used in combination with data assimilation of satellite Aerosol Optical Depth to provide forecasts and re-analyses of atmospheric composition and boundary conditions for regional air quality models. The shift from the current operational aerosol scheme in C-IFS over to GLOMAP-mode is scheduled to take place during 2018.
The GLOMAP-mode code allows several alternative "aerosol configurations" to be run using the same set of FORTRAN subroutines.
In section 12 of the UKCA UMDP, Table 18 shows the standard configuration for GLOMAP in both of these other 2 host model frameworks (TOMCAT and C-IFS-GLOMAP).
In this 7-mode configuration (known as setup 8), GLOMAP runs with all 7 modes activated, with 2 being pure dust modes and the other 5 containing mixtures of different aerosol components (sulphate, black carbon, organic matter, sea-salt and dust).
With this GLOMAP setup 8, the model runs has 7 number mixing ratios (one for each mode) and has a total of 19 component mass mixing ratios over all the different types.
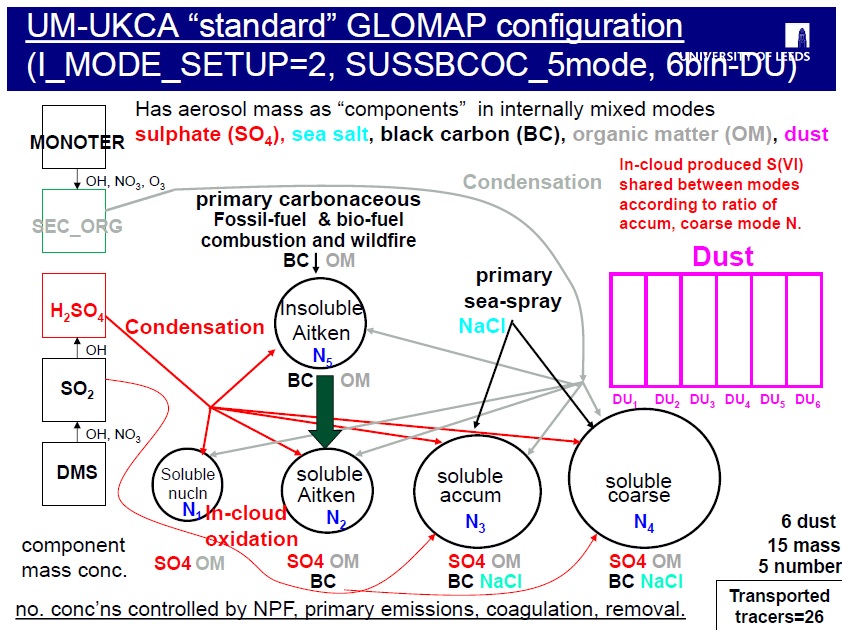
As mentioned in task 10 however, when GLOMAP is run within UM-UKCA, dust is usually handled by the existing 6-bin UM scheme, and GLOMAP is configured to use the "5-mode configuration" (known as setup 2) covering only 4 of the above 5 components (sulphate, black carbon, organic matter and sea-salt).
The scheme can also be further reduced to cover just the sulphate and sea-salt components in 4 modes (known as setup 1), which was the initial configuration used for interactive stratospheric aerosol simulations (e.g. Dhomse et al., 2014).
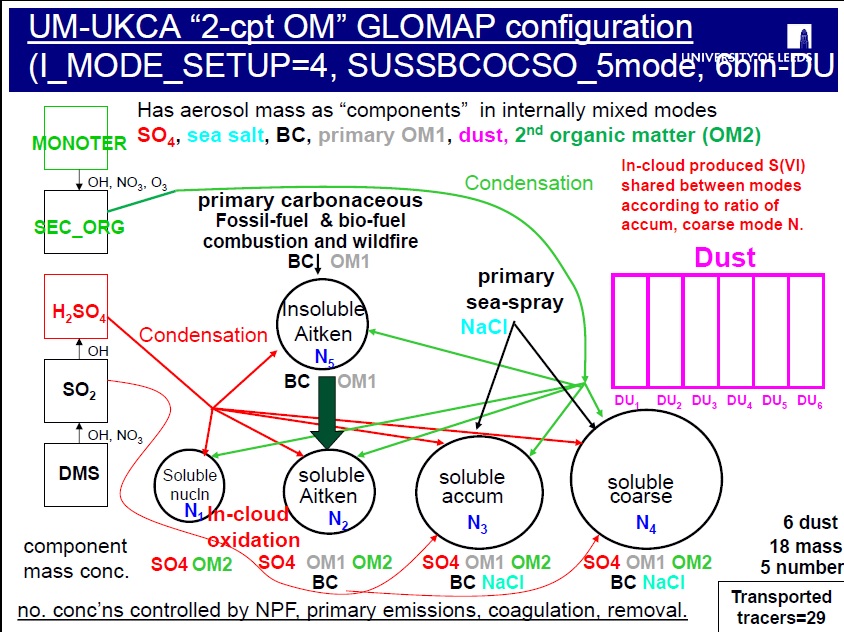
The GLOMAP scheme has been run in the TOMCAT CTM in a range of configurations, including one specifically designed to track two separate components for organic matter (OM), with the I_MODE_SETUP=4 configuration designed to have one component track the primary OM in each mode and a separate OM component to track secondary organic matter in each mode.
Section 12.2 of the UMDP has a more detailed explaination of these configurations with Table 19 showing how these 4 different GLOMAP-mode setups map onto the model tracers.
In this task you will take a copy of the standard UKCA job (which uses GLOMAP-mode setup 2, MS2) and change it to use GLOMAP-mode setup 4 (MS4) to track two separate organic matter (OM) components rather than the usual 1. With the 2-component OC configuration, the model tracks primary (emitted) organic carbon in the usual OM component and secondary organic matter (formed following oxidation in the atmosphere) separately in a 2nd OM component.
Task 11.1: Understand how the GLOMAP aerosol module tracks aerosol species and modes
TASK 11.1: Read section 14 (page 51) of the vn10.4 UKCA Unified Model Documentation Paper (UMDP084) and refer to Tables 28, 29 on pages 53.
Task 11.2: Run a copy of the standard UKCA job which tracks two OM components in the GLOMAP modes
Take a copy of the suite you generated from Task 10 with the GLOMAP AOD diagnostics added. This was your copy of the standard ARCHER v10.4 UKCA tutorial suite (u-ai071). The task is to change the aerosol settings from the standard GLOMAP aerosol configuration (setup 2) to instead use the 2-component OM configuration (setup 4).
To run the 2-component GLOMAP configuration, you will need to add in an additional branch to the suite (which supports that) and also make change to the Rose GUI metadata for the suite, so that an extra option for I_MODE_SETUP=4 is made available within the "GLOMAP configuration" selection-button.
First, add in the extra branch. Go to the fcm_make_um app (4th from the top) and then click on "env" then "Sources" which then shows you the 4 branches that are already used in the UKCA tutorial job. These 4 branches are to enable additional settings over and above what was present in the UM trunk code at v10.4.
To add in the extra branch click the plus sign and then paste the following into the new entry that appears after you click the plus:
branches/dev/grahammann/vn10.4_support_GLOMAPsetup4_for_2cptOM@32315
This is telling to the model that, when the source-code is built, it needs to pick up the code-changes from revision 32315 of the branch I put together to enable UM-UKCA to correctly align with the code that is present in GLOMAP for setup 4 (and adds a few bits that were missing in the v10.4 trunk).
Next we'll need to change the GLOMAP setup. In the Rose GUI, you need to open the UM app (click on the right-hand-pointing triangle at the bottom next to "um") and then similarly click on "namelist" then "UM Science settings" and finally you can then select "Section 34: UKCA" to view which options have been selected in this particular UKCA tutorial job.
There are a long set of different switches which specify how this particular UM-UKCA is configured. Scrolling down (a fair way down) you will see the GLOMAP aerosol setting options listed as Set aerosol species and modes. This basically specifies the value for i_mode_setup, which controls the GLOMAP aerosol configuration (as explained above). In the tutorial job you can see I_MODE_SETUP has been set to 2. That corresponds to the "standard GLOMAP setup" explained and illustrated above (sulphate, sea-salt, black carbon and one organic carbon component in 5 modes).
You see also that there are alternative options there that can change the aerosol setup to 1 (sulphate and sea-salt in 4 modes), 6 (dust-only in 2 modes) and 8 (sulphate, sea-salt, BC, POM and dust in all 7 modes).
In our case, we actually want a different set-up though to any of these standard 4 options -- to activate to use the 2-component organic matter configuration. Of course one needs to know that that configuration is actually available in the code, and although this is explained in the UMDP, it turns out that, at this version, the I_MODE_SETUP=1, 6 and 8 configurations are not supported, and we recommend you do not select those options without corresponding closely with the GLOMAP development team at Leeds.
In order to do this requires to point the Rose GUI to an updated version of the metadata that provides the information for the different panels in the GUI. At the moment, the metadata used in the job tells the UM that the only options available are I_MODE_SETUP=1, 2, 6 and 8. But with the changes implemented in the branch above, it is then possible to also run the model with I_MODE_SETUP=4 (which is the GLOMAP configuration that we want to run with).
The metadata for the UM app within the Rose GUI panel is specified in the first panel selection in the GUI. Instead of clicking on the right-hand-pointing triangle, click on the word um and then you should see a panel with only one entry that says "meta" with the text set as:
/home/grenville/meta/ga7_vn10.4
This is pointing to a rose-meta.conf file that specifies the metadata for the UM app. It is off Grenville Lister's directory because there were things that were decided were required to be added (chosen according to some priorities or requirements) to the functionality of that UM app at that particular time.
In preparation for this task I have updated those Rose selections to additionally allow the user to select the option 4 as well as those 1, 2, 6 and 8.
For this task we need to change the text set in that box to instead be set as:
/home/gmann/meta/ga7_vn10.4_MS4updated
and then click apply. I have updated the rose-meta.conf metadata file to add the required to enable the extra I_MODE_SETUP=4 to be selected in the GUI.
To get an idea of what is different about the rose-meta.conf file, that then allows the user to choose that option, you could use a graphical difference engine like "tkdiff" or "xxdiff" to compare the rose-meta.conf in those two directories.
tkdiff /home/grenville/meta/ga7_vn10.4/rose-meta.conf /home/gmann/meta/ga7_vn10.4_MS4updated/rose-meta.conf
By doing this, you see that the functionality that was added in the file off Grenville Lister's directory was to retain the information in the existing rose-meta.conf, but additional add a new switch to allow the user to scale up or down how much NOx is emitted from lightning sources.
In the rose-meta.conf for the MS4updated version of this, the code does the same to retain the existing settings set in the rose-meta.conf in the branch, and adds in the new lightnight NOx scaling, but then also provides a replacement for the existing I_MODE_SETUP metadata to allow I_MODE_SETUP to take 5 alternative values (1, 2, 4, 6 and 8) instead of the existing 4.
Once you have changed the text to point to that new meta data file, the button-selection in the UKCA panel changes to having those 5 options rather than the previous 4. Now that the new I_MODE_SETUP=4 option is visible in the Rose GUI, proceed to select that I_MODE_SETUP=4 rather than the previous (default) setting of I_MODE_SETUP=2.
The FORTRAN in the model source-code is (with the updated branch added above) able then to act on the new I_MODE_SETUP specied in the Rose GUI, and activate the required extra OM2 tracers (section 34 items 128, 129, 130 and 131) and also does several other things to enable the code to then have the condensation of SEC_ORG go into the SO component mmrs instead of the OM component mmrs as was the case for I_MODE_SETUP=2.
Note that at previous versions of the UM (v7.3, v8.4), to add the extra tracers for the new I_MODE_SETUP required a UMUI hand-edit to edit the file SIZES, setting the values of the array TC_UKCA [which specifies which of the UKCA tracers are switched on (=1) or off (=0)]. By contrast, at v10.4, the control for which tracers should be on or off is done based on what is set in the STASHmaster_A file. There is some code within one of the UKCA routines (tstmask_ukca_mod.F90) that interprets one of the fields set in the STASHmaster_A.
If you have not encountered the so-called "STASHmaster_A" file before, it is basically the file that specifies the properties of all the different prognostic and diagnostic variables that are set to be used by the UM.
The STASHmaster_A file is not easy to understand, and it will not be possible to explain many aspects of what it does -- but suffice it to say that it contains a sequence of 5-line segments for each prognostic or diagnostic variable, with several different fields within it, which conveniently specify a range of information about that particular prognostic/diagnostic's properties (that are interpeted within the FORTRAN code of the model when it runs).
The UKCA prognostic variables can be found in section 34 in items 1-150. All the aerosol optical properties diagnostics (e.g. as explained in Task 10), such as the AOD for each mode, can be found in section 2. (Although strictly speaking this section 2 is principally to store long-wave radiation diagnostics, the aerosol optical properties are also stored here for historical reasons.)
The reason I explain this is that we will refer to the STASHmaster_A file to see the way the main UKCA tracers are specified as on or off for the different I_MODE_SETUPs. And you need to be able to appreciate the context of the GLOMAP tracers that have been activated, in relation to the other variables set for the other parts of the model
We will see that this automatic configuring the tracers is actually done from within one of the fields in the STASHmaster option code entry. The user-prognostic fields required by other parts of the model (e.g. partial volumes for RADAER, CDNC diags for ACTIVATE, aerosol surface area for heterogeneous chemistry) are also able to be conveniently controlled by a similar approach making the extra fields for the different I_MODE_SETUPs available automatically based on what is set in the STASHmaster_A.
Which of the GLOMAP tracers are switched ON for different GLOMAP configurations (values of I_MODE_SETUP) is controlled by a 30-character string on the 3rd line of the STASHmaster_A entry for that variable. This 30-character string is known as an option code. By switched ON I mean it is then set to be a transported tracer and thereby made available during run-time.
If you are familiar with FORTRAN it is worth taking a look at the STASHmaster and also referring to the code where this is done. And for this reason, I recommend at this point that you do an "FCM checkout" (to your local space on PUMA) of the v10.4 branch that was added above.
If you haven't already, you should create an "FCM" directory off your home directory on PUMA. This is where you will store checkouts of branches that you have created (or want to refer to). After cd'ing into that directory, run the following command to "check-out" the code in the branch to a "user working copy" in your ~/FCM directory. The command is:
fcm checkout fcm:um.x_br/dev/grahammann/vn10.4_support_GLOMAPsetup4_for_2cptOM
That checks-out the whole contents of that branch to your ~/FCM/ directory and may take a few minutes to complete (although sometimes it's much quicker).
To look at the STASHmaster_A file, cd into that branch "vn10.4_support_GLOMAPsetup4_for_2cptOM" and then into the following directory within the branch:
cd rose-meta/um-atmos/HEAD/etc/stash/STASHmaster
you will see the STASHmaster_A file.
It's a big file but if you go to line 14028 (e.g. use capital-G in vi) you see section 34 item 101 and on the 3rd line you see the 30-character string of zeros and ones for nucleation mode (soluble) number is set as "100000000000000000000010001011".
The way the model has been coded is to pick up the values of 1 or 0 near the end of this "option code" and use these to specify which tracers are switched on. In short, the nth character from the right-hand side specifies whether the tracer is switched on for I_MODE_SETUP=n.
You see that the 29th and 30th characters are set to "1" which means that the tracer is included for I_MODE_SETUP=1 and I_MODE_SETUP=2. You'll recall that the only other values for I_MODE_SETUP that are allowed to be selected at this model version (in the unchanged base job that is) are I_MODE_SETUP=6 (2mode dust-only) and I_MODE_SETUP=8 (the "full" setup with all 7 modes active). And you see that the 6th-from-the-right is a zero which means that the nucleation mode number tracer is not included in that I_MODE_SETUP=6 configuration (dust-only). But you see that the 8th-from-the-right is set to "1" which means that the tracer is included for I_MODE_SETUP=8 (the "full" set-up with all 7 modes active).
For this task, my branch has additionally activated an extra GLOMAP set-up -- and see that I have edited the STASHmaster to put a 1 also in the 4th character from the end so that that particular tracer is selected to be switched on for the I_MODE_SETUP=4 option which is now available.
That's different from the default specification for this tracer (in the UM trunk at v10.4) which specifies the option code as zero for this column 26:
100000000000000000000010000011
You see in the STASHmaster_A file there are the two sets of organic matter tracers, the first one being called "OC" and the second one called "SO". At some points in the model the tracers are given short-hand names of Ait_SOL_OC (item 106), Acc_SOL_OC (110), Cor_SOL_OC (116), Ait_INS_OC (121) and Nuc_SOL_OC (126) while the second OM tracers have short names Nuc_SOL_SO (128), Ait_SOL_SO (129), Acc_SOL_SO (130) and Cor_SOL_SO (131). But the STASHmaster_A file has their full names -- and you can see that whereas the "OC" tracers are included for both I_MODE_SETUP=2 and I_MODE_SETUP=4, the "SO" tracers are only included for I_MODE_SETUP=4.
The extra tracers for the OM2 mass mixing ratios in each are stored in STASH items 128, 129, 130 and 131 in section 34. You see that these are set to 1 for the SO components in each of the soluble modes and the nucleation soluble OC mmr is no longer required as it has been replaced with SO mmr for that particular mode.
When we changed the Rose GUI to allow the user to select I_MODE_SETUP=4, we did this by pointing the Rose GUI to an updated rose-meta.conf file in a separate directory that enabled the I_MODE_SETUP=4 option. It was this one:
/home/gmann/meta/ga7_vn10.4_MS4updated/rose-meta.conf
The reason I refresh the memory about this is to further explain that, in a sub-directory of that same "ga7_vn10.4_MS4updated" main directory, the "MS4-updated" version of the STASHmaster_A file that I included in the branch is also identically present there (with the I_MODE_SETUP=4 option enabled). This copy of the "MS4-updated" STASHmaster_A file can be found at:
/home/gmann/meta/ga7_vn10.4_MS4updated/etc/stash/STASHmaster
Due to a quirk in this preliminary realisation of the Rose software (which is still in its infancy really for use with the UM), when one makes a change to the STASHmaster_A file, and one wants to use the functionality added to the STASHmaster_A in an actual suite, then one needs to make the change like this in 2 places. In fact, when it comes to running the job, one also has to additionally do a manual copy-in of the STASHmaster_A into a sub-directory of the ~/roses/suite-id/app/um/ part of the suite that one wants to run.
So here we need to also do this manual copy-in of the updated STASHmaster_A (from either the branch or the metadata directory) into a directory with the particular rose suite you are running.
In fact it's more tricky than that because you actually have to create a new directory called file within the app/um/ section of your ~/roses/$SUITE_ID/ file-path and copy-in the updated STASHmaster_A to there. If you don't do this then even though you have updated the STASHmaster_A in the branch, when the model runs it will not use this updated file.
So first you need to cd to ~/roses/$SUITE_ID/app/um and then type mkdir file. Then copy the updated STASHmaster_A file from
/home/gmann/meta/ga7_vn10.4_MS4updated/etc/stash/STASHmaster/
into that new directory you've created called file. Even after you've done that, the changes that have been made to the STASHmaster_A will still not be correctly picked up when the job runs until you have added a STASHmaster_A to alert the Rose GUI that this modified STASHmaster_A needs to be installed when the model runs. It is regrettable that whereas this was done atuomatically at v8.4 it turns out to be rather cumbersome to do it at v10.4.
Anyway, we need to make an additional change in the Rose GUI to alert the model to the fact that the job needs to use a modified STASHmaster_A file. This is done by adding a new environment variable STASHMSTR. Within the um app section of the Rose GUI, go to env and then select Runtime Controls. You see in the right-hand panel it explains this is where environment variables can be set to control behaviour at run time. In our case, this is what we have to do to alert the model that we have changed the STASHmaster_A. It's obviously not ideal that this is not done automatically. But unfortunately that is simply where things are at currently with the new Rose GUI being rolled out before some of the additional functionality is quite ready yet. I expect this will be taken care of at a later UM version but at v10.4 one has to add this extra environment variable to alert the model that things have changed.
So. To add the extra environment variable to alert the model that the STASHmaster_A file has changed. You need to first go to the View panel at the top of the GUI and select View Latent Variables. When you have done that you see that there is in fact an entry for specifying the STASHMSTR environment variable. Click on the plus sign next to that STASHMSTR entry and type the single-character string "." into that panel (that's it just the dot). That specifies to pick up the STASHmaster_A from the file directory.
Next, I advise to look in the directory src/atmosphere/UKCA of the checked-out branch. This is where all the main UKCA code can be found (including the GLOMAP routines) and also the routines for RADAER, ACTIVATE and FAST-J/JX. The reason I mention this is that I think it is worth taking a look at the routine ukca_init.F90 as this gives you some idea of how the code processes these different aerosol configurations and how GLOMAP acts on the different settings of I_MODE_SETUP to specify which modes and components are switched ON and OFF. Looking in ukca_init.F90 you see there is the comment that says "Call appropriate MODE setup routine". You see there that, for each different setting for I_MODE_SETUP, the code calls a different trio of "module-procedures" that together set up everything that is needed to run that particular set-up.
If you already know quite a bit about GLOMAP you could take a look at these by looking in the FORTRAN-90 module ukca_mode_setup.F90
In ukca_init.F90 you see that, for I_MODE_SETUP=2, the code specifies to call the module-procedure "ukca_mode_sussbcoc_5mode" whereas for I_MODE_SETUP=4 it calls "ukca_mode_sussbcocso_5mode". These ukca_mode_xxxx routines specify the main logical variables mode and component which specify which modes are switched on and which components are included in each mode.
The rationale of the GLOMAP setup arrays (set differently in each alternative module-procedure) is that once they are set (along with the associated arrays that specify different properties about each component), the same FORTRAN code can then be run for any supported GLOMAP set-up. For more info about this see the UKCA UMDP (section on GLOMAP aerosol).
The other two setup indices module-procedures specify what is done for processes that exchange from the gas phase to the particle phase (principally condensation of H2SO4 or SEC_ORG into one of the aerosol components in the particle modes). The module-procedures set for example how the condensation routine UKCA_CONDEN should direct the condensation from the gas phase into particular components (by direct I mean which component the gas phase should partition into or out of).
Because of the way the code has been written (reasonably flexibly, but note this is only for tried-and-tested GLOMAP set-ups) once these three module-procedures are set, the model then has all it needs to enable the same code to be run with different module-procedures that specify which modes are to be switched on and which off.
At this point, let's get back to the task in hand. Which is to convert your copy of the UKCA tutorial job to run with the two-component POM setup of GLOMAP (I_MODE_SETUP=4).
Now that your suite points to the updated metadata to enable the I_MODE_SETUP=4 GLOMAP setup, and the Rose GUI is all set with the updated STASHmaster_A (including the environment variable STASHMSTR and the manual copy-in of the STASHmaster_A file) things are pretty much ready to go. You just need to add in the extra daily-mean STASH requests for the additional 4 2nd OM mmr's (the SO tracers) as you did to request the standard OM mmrs in task 10. With both sets of OM mmrs requested to the daily-mean .pk files you will be able to validate that there is the same amount of OM in the two-component OM run as in the standard single-component OM run (with also the aerosol optical properties being equivalent).
To add in the extra STASH requests it's just the same procedure as in Task 10 -- go to Model Input and Output within the um namelist section of the GUI and then STASH Requests and Profiles" and then STASH requests. Click the plus sign next to New in the top right-hand corner of the right-hand panel and then add in the section from section 34 items 128, 129, 130 and 131 for the SO mmrs in each of the four soluble modes. To do this double-click on the item so it is added to the list of STASH requests and then set the profile for these 4 new ones as dom_name as DALLTH (all theta points in the 3D grid), tim_name as TDAYM for daily-mean and use_name as 'UPK' to send these to the .pk files as for the daily-mean OM1 mmrs you already added in Task 10.
Note that whereas at v8.4 the user had to switch to a different version of the RADAER hand-edit raderv2_vn84_ARCHER.ed to allow the 2-component OM configuration of GLOMAP to couple to the UM radiation scheme, at v10.4 this is all taken care of automatically within the code (based on the settings for the required user-prognostics in the STASHmaster_A).
So, as the models runs, the AOD's calculated within RADAER will automatically be including the additional partial volume from the OM2 mmr's (items 128 to 131, which now track only the secondary OM) as well as the partial volumes for the usual OC mmrs (which now only include the primary OM).
Finally, since you have asked the model to run with additional tracers, you also need to specify how these should be initialised.
Go to the Reconfiguration and Ancillary Control panel in the namelist section of the um app within the Rose GUI and select Configure ancils and initialise dump fields. When you select that, the window on the right-hand side should view a whole set of entries which indicate where some of the tracers need a different type of initialisation (not just picking up the values from the dump). These special initialisations are handled via what is referred to as an items namelist.
The information here is hard to de-cipher but the best way to understand it is to look for the column entitled stash_req. That explains which STASH item the panel is specifying needs to be initialised differently. If you click on the stash_req it will re-sort the list to be in ascending order for the STASH requests (put section and items in order).
Basically there are several different ways to initialise tracers (or other model fields) when one runs the model. You can specify that they should be set to zero (source set to 3), that they should pick up values from file or user-ancilliary file (source set to 7), or else have them be set to a constant (non-zero) value (source set to 6). The UM has been coded to allow the user to specify this very flexibly, based on what is set in the GUI (previously UMUI, now Rose GUI). In short, the choice of how the tracers should be initialised is determined by what is set in the column source. If that source column is set to 7 then it means the tracer is to be initiliaised (in an NRUN) by values set in a user-ancilliary file (the name of which is specified in the ancil filename column). For example you see that many of the UKCA tracers are set to be initilised from a pre-existing user-ancillary file -- that is set by the environment variable CHEM_INIT.
Here, we want to initialise the 4 extra OM2 mmrs to be set to zero at the start of the NRUN so we need to add those in and set them to have the source column to be set to 3, for initialising the tracer to zero. To add the extra initialisations you need to right-click in the right-hand window and select Add section. You will then get a new row to the column given the index 1. Then right-click again and select view namelist items. This then sets the different items. You need to add all 4 to this one new section by, for each one, clicking on section and setting this to section 34 and selecting first 128, then 129, then 130 and then 131 -- clicking on the plus sign after the first one and selecting Set to zero from the source section (you only need to do that once). When you add the 2nd one you'll find you need to use the cursor keys due to a bug in the GUI. After clicking Save there will then be an extra line in that list of special initialisations which nows list those extra 4 SO mmrs as being initialised to zero.
That's it! With those changes you should be able now to proceed and submit the job as in Task 10.
There is more info in the UMDP on initialising variables in UMDP302:
https://code.metoffice.gov.uk/doc/um/latest/papers/umdp_302.pdf (password required)
Worked Solution
The worked solution to this task can be found in job u-ai897.
Sample output can be found on ARCHER in the directory
/work/n02/n02/ukca/Tutorial/vn10.4/sample_output/Task11.3
Your .pk file should now contain the following fields
0 : 192 144 6 1 Unspecified 1 : 192 144 6 1 Unspecified 2 : 192 144 6 1 Unspecified 3 : 192 144 6 1 Unspecified 4 : 192 144 6 1 Unspecified 5 : 192 144 6 1 Unspecified 6 : 192 144 6 1 Unspecified 7 : 192 144 6 1 Unspecified 8 : 192 144 6 1 Unspecified 9 : 192 144 6 1 Unspecified 10 : 192 144 6 1 Unspecified 11 : 192 144 6 1 Unspecified 12 : 192 144 85 1 AITKEN MODE (SOLUBLE) OC MMR 13 : 192 144 85 1 ACCUMULATION MODE (SOL) OC MMR 14 : 192 144 85 1 COARSE MODE (SOLUBLE) OC MMR 15 : 192 144 85 1 AITKEN MODE (INSOLUBLE) OC MMR 16 : 192 144 85 1 NUCLEATION MODE (SOLUBLE) OC2 MMR 17 : 192 144 85 1 AITKEN MODE (SOLUBLE) OC2 MMR 18 : 192 144 85 1 ACCUMULATION MODE (SOLUBLE) OC2 MMR 19 : 192 144 85 1 COARSE MODE (SOLUBLE) OC2 MMR
With the exception of the STASHmaster_A file changes, the following Rose edits are required:
Index: app/um/rose-app.conf
===================================================================
--- app/um/rose-app.conf (revision 28536)
+++ app/um/rose-app.conf (revision 28574)
@@ -1,4 +1,4 @@
-meta=/home/grenville/meta/ga7_vn10.4
+meta=/home/gmann/meta/ga7_vn10.4_MS4updated
[command]
default=um-atmos
@@ -23,6 +23,7 @@
RECON_KEEP_MPP_STDOUT=true
RECON_STDOUT_FILE=pe_output/${RUNID}.fort6.pe
SPECTRAL_FILE_DIR=$UMDIR/vn$VN/ctldata/spectral/ga7
+STASHMSTR=.
UM_THREAD_LEVEL=MULTIPLE
VN=10.4
@@ -2083,6 +2084,17 @@
user_prog_ancil_stash_req=34060
!!user_prog_rconst=1.000000e-12
+[namelist:items(9fcddcd3)]
+ancilfilename=
+domain=1
+!!interval=0
+!!period=1
+source=3
+stash_req=34131
+update_anc=.false.
+!!user_prog_ancil_stash_req=0
+!!user_prog_rconst=0
+
[namelist:items(a01f7039)]
ancilfilename='$CHEM_INIT_FILE'
domain=1
@@ -2171,6 +2183,17 @@
user_prog_ancil_stash_req=34150
!!user_prog_rconst=0.0
+[namelist:items(aebf9338)]
+ancilfilename=
+domain=1
+!!interval=0
+!!period=1
+source=3
+stash_req=34130
+update_anc=.false.
+!!user_prog_ancil_stash_req=0
+!!user_prog_rconst=0
+
[namelist:items(af3c6568)]
ancilfilename=
domain=1
@@ -2281,6 +2304,28 @@
!!user_prog_ancil_stash_req=
user_prog_rconst=1.000000e-18
+[namelist:items(d050f041)]
+ancilfilename=
+domain=1
+!!interval=0
+!!period=1
+source=3
+stash_req=34128
+update_anc=.false.
+!!user_prog_ancil_stash_req=0
+!!user_prog_rconst=0
+
+[namelist:items(d13ecc0f)]
+ancilfilename=
+domain=1
+!!interval=0
+!!period=1
+source=3
+stash_req=34129
+update_anc=.false.
+!!user_prog_ancil_stash_req=0
+!!user_prog_rconst=0
+
[namelist:items(dc674670)]
ancilfilename='$CHEM_INIT_FILE'
domain=1
@@ -3957,7 +4002,7 @@
file_strat_aer='Sulfate_SAD_SPARC_Background.asc'
!!i_mode_bln_param_method=0
i_mode_nzts=15
-i_mode_setup=2
+i_mode_setup=4
!!i_so2_hi_level=8
i_ukca_chem=51
i_ukca_conserve_method=2
@@ -4110,6 +4155,14 @@
tim_name='TDMPMN'
use_name='UPMEAN'
+[namelist:streq(25699046)]
+dom_name='DALLTH'
+isec=34
+item=130
+package=
+tim_name='TDAYM'
+use_name='UPK'
+
[namelist:streq(26025673)]
dom_name='DALLTH'
isec=34
@@ -5870,6 +5923,14 @@
tim_name='TALLTS'
use_name='UPUKCA'
+[namelist:streq(45fe4ef4)]
+dom_name='DALLTH'
+isec=34
+item=131
+package=
+tim_name='TDAYM'
+use_name='UPK'
+
[namelist:streq(463b0360)]
dom_name='DALLTH'
isec=34
@@ -6206,6 +6267,14 @@
tim_name='TDMPMN'
use_name='UPMEAN'
+[namelist:streq(51db0630)]
+dom_name='DALLTH'
+isec=34
+item=128
+package=
+tim_name='TDAYM'
+use_name='UPK'
+
[namelist:streq(52a891d5)]
dom_name='DALLTH'
isec=34
@@ -7750,6 +7819,14 @@
tim_name='T6HMONM'
use_name='UPG'
+[namelist:streq(957e3213)]
+dom_name='DALLTH'
+isec=34
+item=129
+package=
+tim_name='TDAYM'
+use_name='UPK'
+
[namelist:streq(95d040d8)]
dom_name='DP17'
isec=30
Index: app/fcm_make_um/rose-app.conf
===================================================================
--- app/fcm_make_um/rose-app.conf (revision 28536)
+++ app/fcm_make_um/rose-app.conf (revision 28574)
@@ -45,4 +45,4 @@
stash_version=1A
timer_version=3A
um_rev=vn10.4
-um_sources=branches/dev/mohitdalvi/vn10.4_scale_lightning_nox@19623 branches/dev/stevenhardiman/vn10.4_ukca_tropopause_amendment@19627 branches/dev/alistairsellar/vn10.4_no_expvolc_so2@19808 branches/dev/marcuskoehler/vn10.4_ukca_fix_glomap_climatol_surfarea@24038
+um_sources=branches/dev/mohitdalvi/vn10.4_scale_lightning_nox@19623 branches/dev/stevenhardiman/vn10.4_ukca_tropopause_amendment@19627 branches/dev/alistairsellar/vn10.4_no_expvolc_so2@19808 branches/dev/marcuskoehler/vn10.4_ukca_fix_glomap_climatol_surfarea@24038 branches/dev/grahammann/vn10.4_support_GLOMAPsetup4_for_2cptOM@32315
These changes (along with the STASHmaster_A file changes) can be found in the file
/home/ukca/Tutorial/worked_solutions/Task11.2/task11.2.rose.diff
on PUMA.
Task 11.3 Examine the simulated total organic carbon in the original and two-cpt OM configurations
In the above Task 11.2 you ran a 2-component OM version of the UKCA tutorial
job (u-ai071), which began from the updated version from
Task 10 where the AOD diags and the extra STASH requests for OM1 mmrs, had
already been added.
You can also refer to the worked solution u-ai897 which I have
configured in this way.
See that u-ai830 was the same as the UKCA tutorial job
(with the STASH items from Task 10 added) and then u-ai897
adds in the required for this Task.
So by now you should have equivalent standard (as u-ai830) and
2-component OM (as u-ai897) versions of the UKCA tutorial job.
In these jobs you have requested numerous daily-mean fields to be output in
the .pk files.
So in your /work/n02/n02/ directory on ARCHER you should have jobida.pk19991201 files for your standard and 2-component OM jobs.
Included in the extra STASH requests are the mass mixing ratios of OM (the standard organic component) and SO (the 2nd organic component) in each mode.
The OM mmrs are STASH section 34, items 126 (nucleation mode), 106 (Aitken-soluble), 110 (accumn-soluble), 116 (coarse-soluble) and 121 (Aitken-insoluble).
The SO mmrs are STASH section 34, items 128 (nucleation mode), 129 (Aitken-soluble), 130 (accumn-soluble), 131 (coarse-soluble).
Note that when running with the two-component OM configuration (I_MODE_SETUP=4), the OM mmr in the nucleation mode (34-126) will always be zero, and for that reason the option code in the 4th-from-the-right column in the STASHmaster_A has been set to zero. As a consequence, there will be no 34-126 code available in the .pk file when you run the job.
These STASH item numbers and the details of the standard and 2-component GLOMAP configurations can be found in the UKCA UMDP section 12 Tables 19 and 20.
Note also that there is no SO in the Aitken-insoluble mode as this contains only primary carbonaceous particles. Any SO or OM condensing onto the particles in the insoluble modes is immediately transferred over to the corresponding soluble mode following the "condensation-ageing" approach used by the model. This OM or SO condensing onto the insoluble particles is a kind of "coating" for the particles making the particles hygroscopic/soluble.
You could also try adding STASH requests for the mmr of the gas phase species MONOTER and SEC_ORG (STASH section 34, items 91 and 92).
As an example I have put here a link to a pdf ![]() OMcomparison
OMcomparison ![]() showing global maps comparing surface OM fields between the reference single-OM-component suite
showing global maps comparing surface OM fields between the reference single-OM-component suite
u-ai830 (top-left, labelled as ai830) and
u-ai895 (top-right, labelled as ai895).
Page 1 of the pdf compares the "total POM mmr" at the surface which is the total particulate organic matter (POM) summing up the mass of OM and SO in each mode.
Pages 2 and 3 show comparisons of "total POM1 mmr" and "total POM2 mmr" which are the sum of the 1st and 2nd organic component over all the modes.
You can see from the example that the "total POM2 mmr" in u-ai830
is zero everywhere. That's because in this job GLOMAP has the standard configuration
with just one organic component.
By contrast the "total POM2 mmr" for u-ai895 has considerable
concentrations in vegetated continental regions.
In this "I_MODE_SETUP=4" configuration, the "SEC_ORG" species (which contains the
secondary organics from monoterpene oxidation) condenses into the "SO" component,
whereas in u-ai830 SEC_ORG condenses into the "OC" component.
The bottom left panel on each page shows a global map of the ratio of the field for the two model runs. One can use this kind of approach to track the fraction of the OM that is biogenic and anthropogenic.
Note however that we initialised the SO mmr's to zero at the start of the 1-day run. So the OC1 mmrs will be spinning down and the SO mmrs will be spinning up. The daily-mean values are averaging over values on each the 1-hour timesteps over which the UKCA chemistry and aerosol processes are integrated. So although the ratio shown in the bottom-left on page 2 is indicative of the biogenic fraction it should be treated with caution as the fields will not have spun-up/down yet.
This task illustrates how one can separate out the aerosol mass from different sources and track them separately via a different aerosol component.
One could also introduce a 2nd gas phase species like "SEC_ORG" to track different types of SOA. For example one could configure the model so that such a 2nd "SEC_ORG2" species held semi-volatile oxidised organic species with only the very low volatility oxidised organics held in the usual "SEC_ORG" species.
Page 4 also confirms there is virtually no difference in simulated AOD between the job that track both types of OM in one component (u-ai830) and the job that has separate components to track primary and secondary OM (u-ai897). The very slight difference apparent in some remote regions (e.g. close to Patagonia), is likely due to differences associated with the so-called MDTfix code which protects against artefacts in simulated particle size introduced by mass-conservation imposed in the advection scheme, and corrects slightly differently when the two separate sets of OM tracers are used.
Worked Solution
You can use the following convsh script (or Xconv) to extract the required fields to netCDF:
#! /usr/bin/env convsh # This file is part of the UKCA Tutorials: # http://www.ukca.ac.uk/wiki/index.php/UKCA_Chemistry_and_Aerosol_Tutorials_at_vn10.4 # Copyright (C) 2015 University of Cambridge # This is free software: you can redistribute it and/or modify it under the # terms of the GNU Lesser General Public License as published by the Free Software # Foundation, either version 3 of the License, or (at your option) any later # version. # It is distributed in the hope that it will be useful, but WITHOUT ANY # WARRANTY; without even the implied warranty of MERCHANTABILITY or FITNESS FOR A # PARTICULAR PURPOSE. See the GNU Lesser General Public License for more details. # You find a copy of the GNU Lesser General Public License at <http://www.gnu.org/licenses/>. # Written by N. Luke Abraham 2017-01-06 <nla27@cam.ac.uk> # you should change the input file name here readfile 0 ai922a.pk19880901 # You may need to change the field numbers (first column in Xconv or when using list_fields) # You should also change the prepended suite-id # These should match these names: # AITKEN MODE (SOLUBLE) OC MMR writefile netcdf ai922_m01s34i106.nc 12 # ACCUMULATION MODE (SOL) OC MMR writefile netcdf ai922_m01s34i110.nc 13 # COARSE MODE (SOLUBLE) OC MMR writefile netcdf ai922_m01s34i116.nc 14 # AITKEN MODE (INSOLUBLE) OC MMR writefile netcdf ai922_m01s34i121.nc 15 # NUCLEATION MODE (SOLUBLE) OC2 MMR writefile netcdf ai922_m01s34i128.nc 16 # AITKEN MODE (SOLUBLE) OC2 MMR writefile netcdf ai922_m01s34i129.nc 17 # ACCUMULATION MODE (SOLUBLE) OC2 MMR writefile netcdf ai922_m01s34i130.nc 18 # COARSE MODE (SOLUBLE) OC2 MMR writefile netcdf ai922_m01s34i131.nc 19
You can then use cdo to sum the fields together:
cdo add ai922_m01s34i106.nc -add ai922_m01s34i110.nc -add ai922_m01s34i116.nc ai922_m01s34i121.nc ai922_OMmmr_cdo.nc cdo add ai922_m01s34i128.nc -add ai922_m01s34i129.nc -add ai922_m01s34i130.nc ai922_m01s34i131.nc ai922_SOmmr_cdo.nc
You can also use Iris:
#!/usr/bin/env python # This file is part of the UKCA Tutorials: # http://www.ukca.ac.uk/wiki/index.php/UKCA_Chemistry_and_Aerosol_Tutorials_at_vn10.4 # Copyright (C) 2015 University of Cambridge # This is free software: you can redistribute it and/or modify it under the # terms of the GNU Lesser General Public License as published by the Free Software # Foundation, either version 3 of the License, or (at your option) any later # version. # It is distributed in the hope that it will be useful, but WITHOUT ANY # WARRANTY; without even the implied warranty of MERCHANTABILITY or FITNESS FOR A # PARTICULAR PURPOSE. See the GNU Lesser General Public License for more details. # You find a copy of the GNU Lesser General Public License at <http://www.gnu.org/licenses/>. # Written by N. Luke Abraham 2017-01-06 <nla27@cam.ac.uk> # preamble import iris # load all OM OC components om=iris.load('./ai922a.pk19880901', [iris.AttributeConstraint(STASH='m01s34i106'), iris.AttributeConstraint(STASH='m01s34i110'), iris.AttributeConstraint(STASH='m01s34i116'), iris.AttributeConstraint(STASH='m01s34i121')]) # load all SO OC components so=iris.load('./ai922a.pk19880901', [iris.AttributeConstraint(STASH='m01s34i128'), iris.AttributeConstraint(STASH='m01s34i129'), iris.AttributeConstraint(STASH='m01s34i130'), iris.AttributeConstraint(STASH='m01s34i131')]) # make cube to store total OM omsum=om[0].copy() # make cube to store total SO sosum=so[0].copy() # add-up components omsum.data=om[0].data+om[1].data+om[2].data+om[3].data sosum.data=so[0].data+so[1].data+so[2].data+so[3].data # rename omsum.rename('mass_fraction_of_total_particulate_organic_matter_in_air') sosum.rename('mass_fraction_of_total_particulate_organic_matter_in_air') # remove unlimited dimension when writing to netCDF iris.FUTURE.netcdf_no_unlimited=True # output to netCDF iris.save(omsum,'ai922_OMmmr_iris.nc',netcdf_format='NETCDF3_CLASSIC') iris.save(sosum,'ai922_SOmmr_iris.nc',netcdf_format='NETCDF3_CLASSIC')
The calc_AOD.py file provided for the previous Tutorial will also work in exactly the same way here.
These scripts and sample output can be found in the directory
/work/n02/n02/ukca/Tutorial/vn10.4/sample_output/Task11.3
on ARCHER.
Written by Graham Mann 2016