UKCA Chemistry and Aerosol vn10.4 Tutorial 4
UKCA Chemistry and Aerosol Tutorials at vn10.4
What you will do in this tutorial
In this tutorial you will learn how to make the required changes in Rose and in UKCA to include new chemical tracers. As you learn how to do this you will add two new tracers to the UKCA tutorial suite you have already been running.
Task 4.1: Add new tracers into Rose and UKCA
TASK4.1: Add in two new tracers in to slots 64 and 65 for the StratTrop chemistry scheme. The tracer in slot 64 will be called ALICE and the tracer in slot 65 will be called BOB and initialise these tracers to 1.000000e-12. You should also output these two tracers through the UPK stream in STASH as daily means. In UKCA, you should set the conversion factor for each of these to 1.0.
Make a branch
As UKCA is a framework, it must have the infrastructure around it to allow it to work, with the infrastructure being the UM itself. In this tutorial you will add in two new chemical tracers. In this past this was done in two steps, but in Rose this must be done in a single step from vn10.6 onwards. Even though these tutorials are at vn10.4, we will follow the same steps as for vn10.6.
For this tutorial you will need to make a new branch in the usual way. You should do this now.
You will have learned how to make UM branches in the NCAS-CMS FCM Tutorial. However, now you should use um.x_tr as the source of your branch.
For example, first you should make a ticket on the Met Office SRS Trac pages (login required) and then make the branch like so
fcm branch-create --type dev -k ticket_number your_branch_name fcm:um.x_tr@vn10.4
before checking-out your branch by
fcm checkout fcm:um.x_br/dev/userid/vn10.4_your_branch_name
More information on FCM can be found at the NCAS-CMS FCM pages or the FCM User Guide
Metadata
In Rose, the GUI appearance is controlled by metadata, and it is possible to edit this metadata to add new variables to panels. We will not do this in this tutorial, but you will need to change where Rose looks for metadata, as the STASHmaster_A is also metadata.
vn10.4
At vn10.4, the um metadata associated with your suite can be found by clicking on the um menu entry. In the base suite, this is set to /home/grenville/meta/ga7_vn10.4. However, you will also need to include the metadata changes here. To do this, cd into your branch and merge-in the branch fcm:um.x_br/dev/lukeabraham/vn10.4_ga7_meta
fcm merge fcm:um.x_br/dev/lukeabraham/vn10.4_ga7_meta
and then
fcm commit
vn10.6+
At vn10.6, this is not necessary, as the additional metadata for GA7.0 is already in the trunk.
Using your new metadata within Rose
To pick-up the metadata changes, you should edit the meta path in um to point to
um-atmos/HEAD
From now on, when you want to open your Rose suite for editing, instead of doing it through rosie, you should instead open the job for editing by going to your $HOME/roses/[SUITE-ID] directory, and opening it using the command
rose edit -M /path/to/vn10.4_your_branch_name/rose-meta
If you don't do this, the STASHmaster changes will not be picked up, and you won't be able to select the tracers.
Rose Changes
Include your branch for code changes
Although we haven't made any code changes yet, you will (at some point) need to include your branch in Rose so that the UKCA code changes can be picked-up and compiled. To do this go to fcm_make_um env Sources and add a new branch by clicking the plus symbol in the um_sources section. You should then put
/path/to/vn10.4_your_branch_name
if you wish to run from a working copy, and
branches/dev/[your MOSRS username]/vn10.4_your_branch_name
if you want to run from the repository. You can specify specific revision numbers by putting @REV at the end of this line (where REV is the revision number, e.g. 12345 etc.).
STASHmaster_A
In Rose-based jobs, adding new UKCA tracers is done through editing the STASHmaster_A file directly. Take a look at the STASHmaster_A file, found in
/path/to/vn10.4_your_branch_name/rose-meta/um-atmos/HEAD/etc/stash/STASHmaster/STASHmaster_A
This is a large file, as it contains all the possible STASH items. Scroll down the file until you get to section 34, and you will see how the tracers are defined. The key entries to consider are highlighted in red. All entried are fixed-width, so when editing the file, ensure not to change the spacing, otherwise you will get errors.
#=============================================================================== # Section 34 UKCA Chemistry #=============================================================================== # 1| 1 | 34 | 1 |O3 MASS MIXING RATIO AFTER TIMESTEP | 2| 2 | 0 | 1 | 1 | 2 | 40 | 11 | 0 | 0 | 0 | 0 | 3| 000000000000000000000000111110 | 00000000000000000001 | 1 | 4| 1 | 0 | -99 -99 -99 -99 -99 -99 -99 -99 -99 -99 | 5| 0 | 2101 | 0 | 65 | 0 | 0 | 0 | 0 | 0 | #
The meaning of these entries is:
First Line:
- 34: The STASH section, in this case Section 34, UKCA prognostics.
- 1: The STASH item, in this case, the first item in s34, ozone.
- O3 MASS MIXING RATIO AFTER TIMESTEP: The name of the field in the STASHmaster file. This can be anything descriptive, but it isn't read by the model, and is only considered in Rose (or e.g. Xconv).
Second Line:
- 2: The Space Code, which tells the UM whether the field should be held in the restart dump or not. For prognostics, this should be 2 (which means that they should be stored in the restart dump), but for diagnostics that are only passed to output files, this should be 0.
Third Line:
- 000000000000000000000000111110: These are Options Codes which determine which scheme the STASH field (i.e. tracer in this case) are valid for. These are counted from the right, n1 to n30, and for UKCA only the first 8 are used. These have the following meaning:
- n1: Age-of-air tracer (ageair) or aerosol tracers
- n2: Standard Tropospheric Chemistry scheme using the Backward Euler solver (trop)
- n3: Regional Air Quality Chemistry scheme using the Backward Euler solver (raq)
- n4: Tropospheric Chemistry with Isoprene scheme using the Newton-Raphson solver (tropisop)
- n5: Stratospheric-Tropospheric Chemistry using the Newton-Raphson solver (strattrop, also know as CheST)
- n6: Standard Stratospheric Chemistry using the Newton-Raphson solver (strat)
- n7: Offline-oxidants scheme (to drive GLOMAP-mode) using the Newton-Raphson solver (offline)
- n8: Offline-oxidants scheme (to drive GLOMAP-mode) using the Newton-Raphson solver (offline_be)
Therefore, looking at the option code for ozone (s34i001), it is valid for the trop, raq, tropisop, strattrop, and strat chemistry schemes.
Fifth Line:
- 2101: This is the PP Field Code and should be unique to each tracer. Here these begin counting at 2101 (s34i001) and end at 2250 (s34i150) for vn10.4, or 2356 (s34i256) for vn10.6, as the number of tracers has been increased.
So to make a STASH entry for a single tracer called ALICE in slot s34i064 that is only valid in the StratTrop scheme, the STASHmaster_A entry would look like
# 1| 1 | 34 | 64 |ALICE MASS MIXING RATIO AFTER TSTEP | 2| 2 | 0 | 1 | 1 | 2 | 40 | 11 | 0 | 0 | 0 | 0 | 3| 000000000000000000000000010000 | 00000000000000000001 | 1 | 4| 1 | 0 | -99 -99 -99 -99 -99 -99 -99 -99 -99 -99 | 5| 0 | 2164 | 0 | 65 | 0 | 0 | 0 | 0 | 0 | #
Remember: You need to add 2 tracers, ALICE in s34i064 and BOB in s34i065.
You should make these changes to STASHmaster_A, and then fcm commit these changes. This is important, otherwise ARCHER will not be able to see the changes you have made once you have included your new STASHmaster_A file in Rose.
Section 51
All STASH entries in Section 34 need a corresponding pressure-level field to be added to section 51. The equivalent STASHmaster_A entry for s51 ozone is:
# #=============================================================================== # Section 51 UKCA Chemistry on pressure levels #=============================================================================== # 1| 1 | 51 | 1 |O3 MASS MIXING RATIO ON PRESS LEVS | 2| 0 | 0 | 1 | 1 | 3 | 1 | 2 | 0 | 0 | 0 | 1 | 3| 000000000000000000000000111110 | 00000000000000000001 | 3 | 4| 1 | 0 | -99 -99 -99 -99 -99 -99 -99 -99 -99 -99 | 5| 0 | 2101 | 0 | 8 | 0 | 0 | 0 | 0 | 0 | #
You will notice that there are a number of changes to various codes, and this is to do with defining the grid that the diagnostic is valid on, whether it can be held in the dump, etc. For more information on what these codes mean, please see Appendix C in UMDP C4, which can be obtained from the SRS here (password required). When making a new entry, the easiest thing to do is copy an existing entry and make the required changes to the item, name, PP field code, and option codes.
Remember: You need to add 2 tracers, ALICE in s34i064 and BOB in s34i065.
Help text
You should also add appropriate help text into the vn10.4_your_branch_name/rose-meta/um-atmos/HEAD/etc/stash/STASHmaster/STASHmaster-meta.conf file, e.g.
[stashmaster:code(34064)] description=ALICE MASS MIXING RATIO AFTER TIMESTEP help=UKCA Tutiorial Tracer ALICE Mass Mixing Ratio in kg/kg(Air)
and
[stashmaster:code(51064)]
description=ALICE MASS MIX RATIO ON PRESS LEVS
help=UKCA Tutiorial Tracer ALICE Mass Mixing Ratio in kg/kg(Air)
=on pressure levels
Remember: You need to add 2 tracers, ALICE in s34i064 and BOB in s34i065.
Use your new STASHmaster file in Rose
To get Rose to recognise your STASHmaster file, you will need to make a number of changes:
- Point the um metadata to um-atmos/HEAD and edit Rose using
rose edit -M /path/to/vn10.4_your_branch_name/rose-meta(you should already be doing this after following the instructions in the metadata section above. - You need to open the
$HOME/roses/[SUITE-ID]/app/um/rose-app.conffile in a text editor (e.g. vim, emacs, nedit etc.) and add the lineSTASHMSTR=STASHmasterinside the[env]block - You need to open the
$HOME/roses/[SUITE-ID]/rose-suite.conffile in a text editor and add the following lines at the top of the file:
[file:app/um/file/STASHmaster]
source=fcm:um.xm_br/dev/[your MOSRS userid]/vn10.4_your_branch_name/rose-meta/um-atmos/HEAD/etc/stash/STASHmaster@HEAD
- You need to ensure that you have committed your STASHmaster_A changes, otherwise your new tracer(s) will not be seen on ARCHER during the reconfiguration or atmosphere steps.
Initialise your new tracer(s)
Your new tracer(s) is a prognostic variable, as therefore the UM requires prior knowledge of the values it has to be able to run properly. To do this you need to go to um namelist Reconfiguration and Ancillary Control Configure ancils and initialise dump fields. Here you will see a list of fields. You can change the order of these by e.g. STASH number, as this often makes it easier to view.
To add new fields, right click anywhere within the table and click Add new section. This will make a new entry with index 1, if you add another entry, it will be given index 2 etc.. Right-click on this and click on View namelist:items(X) (where X is the index number). This will open a new tab where you can select the STASH section and item for the field you want to initialise using the drop-down menus. Some UKCA tracers need to be given full 3D initial conditions in ancillary file format. However, for this example we will initialise the tracer to a small number, . To do this, select the radio button for Set to a specified constant value and put 1.000000e-12 in the dialog box labelled user_prog_rconst at the bottom of the panel.
You will need to do this for each of the tracers you are adding. When you have finished, you will need to go to the STASH Requests panel (see the STASH tutorial) and run the TidyStashTransform macro to correctly generate the required index value for each tracer initialisation.
At this point, a vn10.4 job (in fact, all Rose jobs prior to vn10.5) will be able to run and output these tracers. However, at vn10.6 the number of tracers was increased to 256, and when doing this extra checks were included that prevented jobs from running with s34 tracers turned on, but without UKCA using them. Because of this we will also include the UKCA changes now as well.
Output your new tracers
To output your tracers, go to the STASH panel and output your tracers as described in the What is STASH? tutorial.
Remember: You need to add 2 tracers, ALICE in s34i064 and BOB in s34i065.
Required UKCA Code Changes
ukca_set_nmspec.F90
Inside UKCA there is a master mapping array that tells the UKCA code which transported species are placed into a particular item number in section 34. At vn10.4 these are contained in the first 150 slots, and at vn10.6 these are in the first 256 slots.
This array is called nm_spec, and it can be found in ukca_set_nmspec.F90. At vn10.4, it looks like this:
nm_spec(1:a_max_ukcavars) = (/ & 'O3 ','NO ','NO3 ','NO2 ','N2O5 ', & 'HO2NO2 ','HONO2 ','H2O2 ','CH4 ','CO ', & !10 'HCHO ','MeOOH ','HONO ','C2H6 ','EtOOH ', & 'MeCHO ','PAN ','C3H8 ','n-PrOOH ','i-PrOOH ', & !20 'EtCHO ','Me2CO ','MeCOCH2OOH','PPAN ','MeONO2 ', & 'O3_S ','C5H8 ','ISOOH ','ISON ','MACR ', & !30 'MACROOH ','MPAN ','HACET ','MGLY ','NALD ', & 'HCOOH ','MeCO3H ','MeCO2H ','H2O ','ISO2 ', & !40 'Cl ','ClO ','Cl2O2 ','OClO ','Br ', & 'BrO ','BrCl ','BrONO2 ','N2O ','HCl ', & !50 'HOCl ','HBr ','HOBr ','ClONO2 ','CFCl3 ', & 'CF2Cl2 ','MeBr ','N ','O(3P) ','MACRO2 ', & !60 'MeCl ','CF2ClBr ','CCl4 ','CF2ClCFCl2','CHF2Cl ', & 'MeCCl3 ','CF3Br ','H2OS ','CH2Br2 ','H2 ', & !70 'DMS ','SO2 ','H2SO4 ','MSA ','DMSO ', & 'NH3 ','CS2 ','COS ','H2S ','H ', & !80 'OH ','HO2 ','MeOO ','EtOO ','MeCO3 ', & 'n-PrOO ','i-PrOO ','EtCO3 ','MeCOCH2OO ','MeOH ', & !90 'Monoterp ','Sec_Org ','SESQUITERP','SO3 ','AROM ', & 'O(3P)_S ','O(1D)_S ','NO2 ','BrO ','HCl ', & !100 'Nuc_SOL_ND','Nuc_SOL_SU','Ait_SOL_ND','Ait_SOL_SU','Ait_SOL_BC', & 'Ait_SOL_OC','Acc_SOL_ND','Acc_SOL_SU','Acc_SOL_BC','Acc_SOL_OC', & !110 'Acc_SOL_SS','Acc_SOL_DU','Cor_SOL_ND','Cor_SOL_SU','Cor_SOL_BC', & 'Cor_SOL_OC','Cor_SOL_SS','Cor_SOL_DU','Ait_INS_ND','Ait_INS_BC', & !120 'Ait_INS_OC','Acc_INS_ND','Acc_INS_DU','Cor_INS_ND','Cor_INS_DU', & 'Nuc_SOL_OC','Ait_SOL_SS','Nuc_SOL_SO','Ait_SOL_SO','Acc_SOL_SO', & !130 'Cor_SOL_SO','Nuc_SOL_NH','Ait_SOL_NH','Acc_SOL_NH','Cor_SOL_NH', & 'Nuc_SOL_NT','Ait_SOL_NT','Acc_SOL_NT','Cor_SOL_NT','XXX ', & !140 'Anth_Prec ','Bio_Prec ','Anth_Cond ','Bio_Cond ','XXX ', & 'XXX ','XXX ','XXX ','PASSIVE O3','AGE OF AIR' & !150 /)
This array is case sensitive and each string is a fixed length of 10 characters, with the whitespace being made up of spaces and not another tpye of white space (e.g. tabs).
When adding new tracers you should not overwrite tracers that are already in use by the chemistry scheme that you are using (e.g. StratTrop etc.), and it is best to also take care and avoid tracers in use by other schemes, if possible. You can see the tracers used by each scheme by looking in the chch_defs array at the top of each ukca_chem_scheme.F90 (e.g. ukca_chem_strattrop.F90 etc.).
Here you should put a new species, 'ALICE ', into tracer slot 64 (current specified as 'CF2ClCFCl2', which is not in use by any scheme at the moment).
Remember: You need to add 2 tracers, ALICE in s34i064 and BOB in s34i065.
ukca_chem_scheme.F90
We will go through changes that need to be made for the StratTrop chemistry scheme (defined in the file ukca_chem_strattrop.F90), but equivalent changes would need to be made for others, e.g. TropIsop or Strat. For this example we will only refer to this scheme.
In this file the list of species used in chemistry is defined by the array chch_defs_strattrop. In fact, there are two copies of this list, chch_defs_strattrop_chem and chch_defs_strattrop_aer. This is due to some slight differences when adding in the required chemistry to drive GLOMAP-mode. The list of species is held in a derived type, of the format:
chch_t( N,'SPECIES ', X,'TYPE ','FAMILY ', D, W, E), &
Where:
Nis a dummy integer. The value doesn't really matter, although it is usually used to number the species in the list.'SPECIES 'is the name of the species, e.g. 'O3 'Xnumber of odd atoms'TYPE 'describes whether the species is a tracer ('TR '), a steady-state species ('SS '), a constant ('CT '), or a constant field ('CF '). For SS, CT, and CF, special code will need to be added.'FAMILY 'is the family that the species belongs to. This field is not currently used.Dis 1 if the species is dry-deposited, and 0 otherwise.Wis 1 if the species is wet-deposited, and 0 otherwise.Eis 1 if the species is emitted, and 0 otherwise. This field is not currently used.
Therefore to add-in the ALICE species, we should insert a line similar to this one
chch_t( 76,'ALICE ', 1,'TR ',' ', 0, 0, 1) &
into chch_defs_strattrop_chem and chch_defs_strattrop_aer, remembering to also increment the size of these arrays
When inserting the tracer into chch_defs_strattrop_aer, some care must be taken if you also want the species to be dry and/or wet deposited at some point, as the arrays that define these are order specific. It is best therefore to insert the tracer into the equivalent place in both chch_defs_strattrop, which will often mean inserting them before the aerosol-chemistry-specific species (i.e. DMS, SO2, H2SO4, MSA, DMSO, NH3, CS2, COS, H2S, SO3, Monoterp, and Sec_Org). If the tracers that you are adding are only going to be a part of the aerosol chemistry used to drive GLOMAP-mode, then it is fine to just edit the chch_defs_strattrop_aer array and put them at the end of the list.
Remember: You need to add 2 tracers, ALICE in s34i064 and BOB in s34i065.
ukca_setd1defs.F90
In ukca_setd1defs.F90 you will find code blocks which set values for various parameters depending on the scheme used. In this example you will need to scroll down to the section which defines the StratTrop (also known as CheST) chemistry, which is located in the Stratospheric Chemistry section and is referenced by using the L_ukca_strattrop logical.
The IF block tests against L_ukca_strattrop and L_ukca_achem (which determines whether or not you require the additional chemistry used to drive the GLOMAP-mode aerosol scheme). You should increment the value of n_chem_tracers by the number of chemical tracers that you are adding in the correct sections of the IF block.
For example, if you only added the tracers to chch_defs_strattrop_aer (and are only interested in using these new tracers with GLOMAP-mode turned on) then you only need to edit the value of n_chem_tracers in the (L_ukca_strattrop .AND. L_ukca_achem) section etc.
If you are adding to a different chemistry scheme then you will need to make these changes accordingly.
Remember: You need to add 2 tracers, ALICE in s34i064 and BOB in s34i065.
ukca_constants.F90
The unit of the tracers is kg(species)/kg(air) (i.e. mass mixing ratio, or mmr), but inside UKCA these species are converted to volume mixing ratio (or vmr). To enable UKCA to do this you will need to add the conversion factor(s) for your new tracer(s) into the code. This is done in the ukca_constants.F90 module.
Open this file and add the required conversion factor(s). The naming convention for these is M_species is the molecular mass of the new species in g/mol, and C_species for the converson factor from vmr to mmr (calculated as M_species/M_air, where M_air=28.97). In actuality, only the C_species listing is always required, although you may need to add the M_species value later if you are emitting into the new species that you are adding.
For example:
REAL, PARAMETER :: C_ALICE = 1.0000
Remember: You need to add 2 tracers, ALICE in s34i064 and BOB in s34i065.
ukca_cspecies.F90
After you have added in the C_species conversion factors, you will need to tell UKCA to use them for your species. To do this you need to edit the ukca_cspecies.F90 module, which contains code which contructs the c_species array of conversion factors for the advected tracers. This contains a subroutine called ukca_calc_cspecies which has a long block of code that you need to edit to add an entry like this:
WHERE (advt == 'ALICE ') c_species = C_ALICE
The advt array is automatically generated by UKCA at run-time from the chch_defs_scheme chemistry scheme definition you edited earlier, so your new tracer(s) will exist within it. You need to add in a new line for each of your tracers which sets the value of the c_species array to your individual C_species parameter. Add the line(s) in at the end of the block.
Remember: You need to add 2 tracers, ALICE in s34i064 and BOB in s34i065.
ukca_setup_chem_mod.F90
The changes made to ukca_setd1defs.F90 told UKCA how many tracers to expect in Section 34. However, you also need to tell UKCA how many new species you have, and how many of them are tracers. While you have added code to ukca_chem_scheme.F90 (e.g. ukca_chem_strattrop.F90), you also need to tell the chemical solver how many species and tracers to expect.
This is done in the routine ukca_setup_chem_mod.F90. Within this routine is a large CASE statement that is used to define various parameters used by the chemical solver. For the StratTrop scheme you should find the section referenced by
CASE (i_ukca_chem_strattrop)
and you should increment the values of jpctr (which defines the number of tracers used in the scheme) and jpspec (which defines the number of species used in the scheme, and also includes steady-state and constant species) by the number of tracers you have added.
Remember: You need to add 2 tracers, ALICE in s34i064 and BOB in s34i065.
Run your suite
Now that you have made the required changes to Rose and in your branch, please run your suite. You may find that it fails on the, fcm_make2_um, recon, or atmos_main jobs. To find the errors and see output, you should go to the
$HOME/[your ARCHER userid]/cylc-run/[SUITE-ID]
directory on ARCHER. Within this there are a number of directories. To see the log files, cd into
log/job/X/[JOB NAME]/NN
(where X will be either 19880901T0000Z on ARCHER, or may be 1 if you are using a vn10.6 suite based on a Rose stem testing job) and view the job.err or job.out files. When the suite successfully runs, the fieldsfile output can usually be found in
$HOME/[your ARCHER userid]/cylc-run/[SUITE-ID]/share/data/History_Data
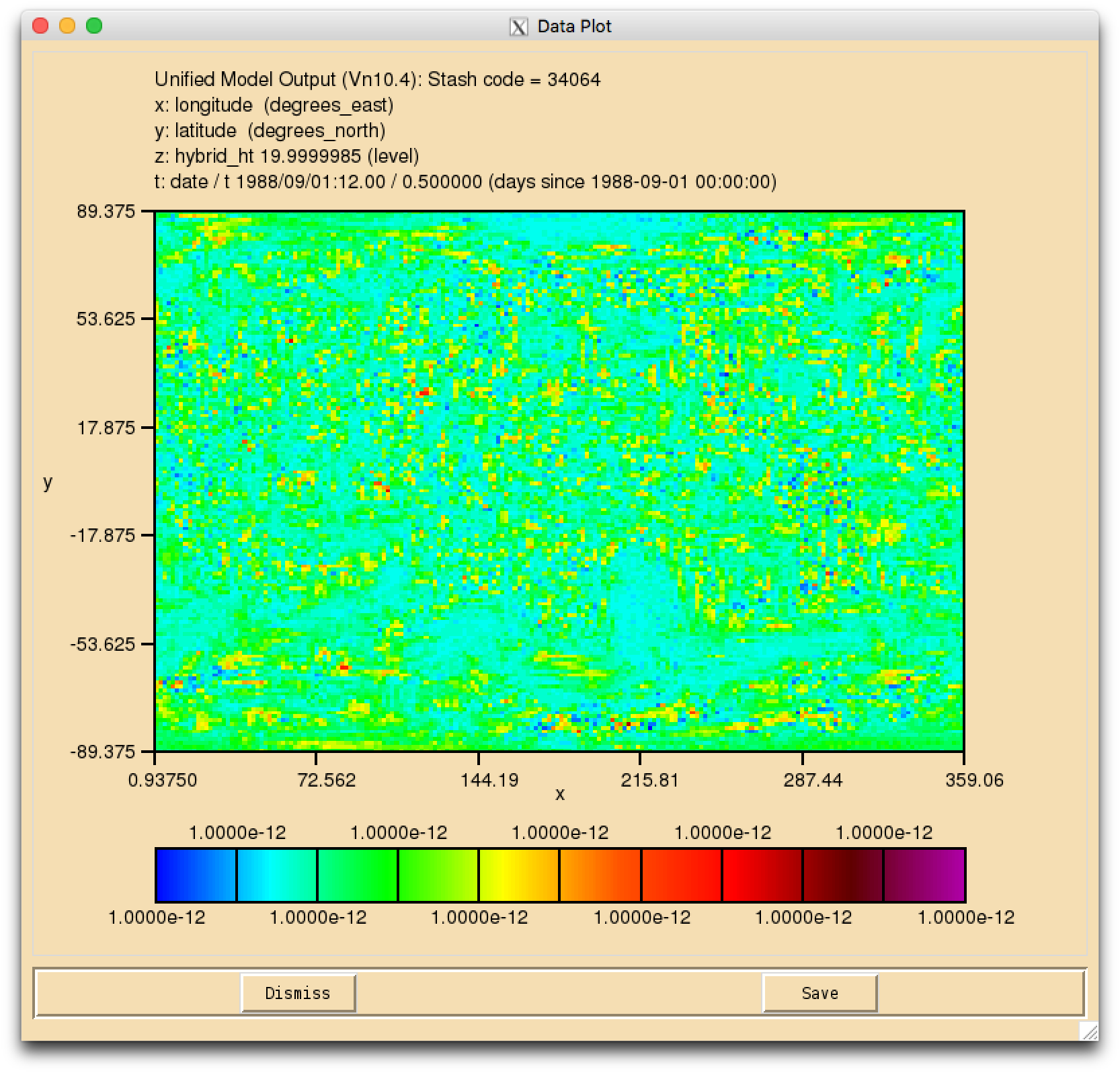
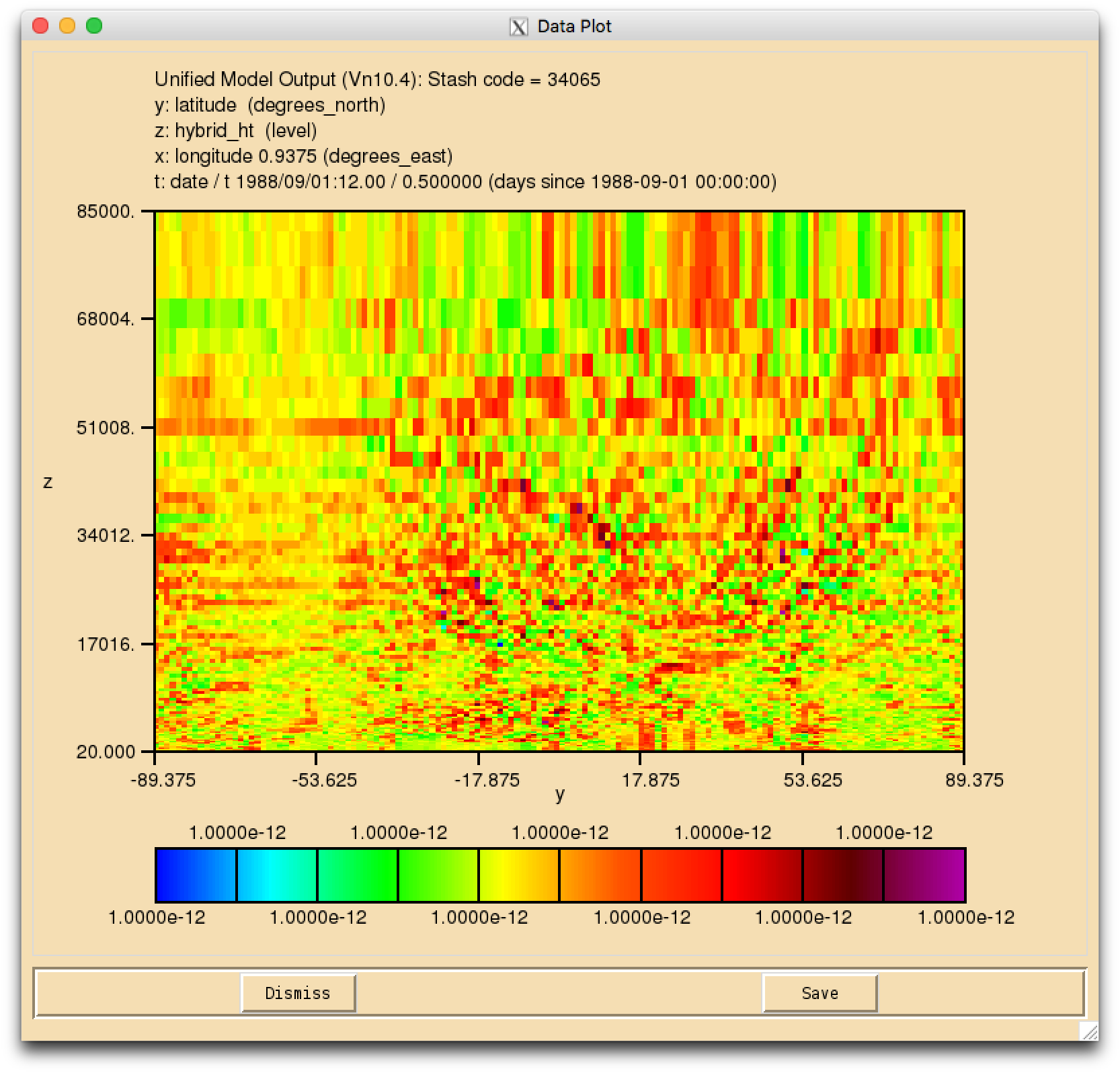
Numerical Noise
When your suite has run successfully, and you view the fields for your new tracer(s) in Xconv, you may find that they have a speckled appearance, rather than being constant. This is due to numerical noise that has been introduced as the tracer is run through the UKCA chemical solver. Once the field is converted to 32-bit this should disappear. It will also cease to become apparent when emissions and reactions are applied to these tracers.
Solution to Task 4.1
You were given the task
- Add in two new tracers in to slots 64 and 65 for the StratTrop chemistry scheme. The tracer in slot 64 will be called ALICE and the tracer in slot 65 will be called BOB and initialise these tracers to
1.000000e-12. You should also output these two tracers through the UPK stream in STASH as daily means. In UKCA, you should set the conversion factor for each of these to 1.0.
For a working Rose suite that has completed this task, please see u-ai084@25815
The specific Rose changes made are:
Index: trunk/app/um/rose-app.conf
===================================================================
--- trunk/app/um/rose-app.conf (revision 25605)
+++ trunk/app/um/rose-app.conf (revision 25815)
@@ -1,4 +1,4 @@
-meta=/home/grenville/meta/ga7_vn10.4
+meta=um-atmos/HEAD
[command]
default=um-atmos
@@ -23,6 +23,7 @@
RECON_KEEP_MPP_STDOUT=true
RECON_STDOUT_FILE=pe_output/${RUNID}.fort6.pe
SPECTRAL_FILE_DIR=$UMDIR/vn$VN/ctldata/spectral/ga7
+STASHMSTR=STASHmaster
UM_THREAD_LEVEL=MULTIPLE
VN=10.4
@@ -1973,6 +1974,17 @@
!!user_prog_ancil_stash_req=
user_prog_rconst=1.000000e-18
+[namelist:items(8dd1e33d)]
+ancilfilename=
+domain=1
+!!interval=0
+!!period=1
+source=6
+stash_req=34064
+update_anc=.false.
+!!user_prog_ancil_stash_req=0
+user_prog_rconst=1.000000e-12
+
[namelist:items(8f0cb76a)]
ancilfilename='$CHEM_INIT_FILE'
domain=1
@@ -2281,6 +2293,17 @@
!!user_prog_ancil_stash_req=
user_prog_rconst=1.000000e-18
+[namelist:items(d9a844ee)]
+ancilfilename=
+domain=1
+!!interval=0
+!!period=1
+source=6
+stash_req=34065
+update_anc=.false.
+!!user_prog_ancil_stash_req=0
+user_prog_rconst=1.000000e-12
+
[namelist:items(dc674670)]
ancilfilename='$CHEM_INIT_FILE'
domain=1
@@ -4574,6 +4597,14 @@
tim_name='TDMPMN'
use_name='UPMEAN'
+[namelist:streq(114ee958)]
+dom_name='DALLTH'
+isec=34
+item=64
+package=
+tim_name='TDAYM'
+use_name='UPK'
+
[namelist:streq(1177a86c)]
dom_name='DIAG'
isec=2
@@ -5838,6 +5869,14 @@
tim_name='TALLTS'
use_name='UPUKCA'
+[namelist:streq(4455fd38)]
+dom_name='DALLTH'
+isec=34
+item=65
+package=
+tim_name='TDAYM'
+use_name='UPK'
+
[namelist:streq(44a86587)]
dom_name='DPFTS'
isec=3
Index: trunk/app/fcm_make_um/rose-app.conf
===================================================================
--- trunk/app/fcm_make_um/rose-app.conf (revision 25605)
+++ trunk/app/fcm_make_um/rose-app.conf (revision 25815)
@@ -45,4 +45,4 @@
stash_version=1A
timer_version=3A
um_rev=vn10.4
-um_sources=branches/dev/mohitdalvi/vn10.4_scale_lightning_nox@19623 branches/dev/stevenhardiman/vn10.4_ukca_tropopause_amendment@19627 branches/dev/alistairsellar/vn10.4_no_expvolc_so2@19808 branches/dev/marcuskoehler/vn10.4_ukca_fix_glomap_climatol_surfarea@24038
+um_sources=branches/dev/mohitdalvi/vn10.4_scale_lightning_nox@19623 branches/dev/stevenhardiman/vn10.4_ukca_tropopause_amendment@19627 branches/dev/alistairsellar/vn10.4_no_expvolc_so2@19808 branches/dev/marcuskoehler/vn10.4_ukca_fix_glomap_climatol_surfarea@24038 branches/dev/lukeabraham/vn10.4_UKCA_Tutorial_Solns@30979
Index: trunk/rose-suite.conf
===================================================================
--- trunk/rose-suite.conf (revision 25605)
+++ trunk/rose-suite.conf (revision 25815)
@@ -1,3 +1,6 @@
+[file:app/um/file/STASHmaster]
+source=fcm:um.xm_br/dev/lukeabraham/vn10.4_UKCA_Tutorial_Solns/rose-meta/um-atmos/HEAD/etc/stash/STASHmaster@HEAD
+
[jinja2:suite.rc]
!!ACCOUNT_USR='foundation'
ANCIL_OPT_KEYS=
For a working UM branch that has completed this task, please see fcm:um.x_br/dev/lukeabraham/vn10.4_UKCA_Tutorial_Solns@30979
The specific UM changes made are:
Index: src/atmosphere/UKCA/ukca_setd1defs.F90
===================================================================
--- src/atmosphere/UKCA/ukca_setd1defs.F90 (revision 30974)
+++ src/atmosphere/UKCA/ukca_setd1defs.F90 (revision 30979)
@@ -309,7 +309,7 @@
(/'NO ','CH4 ','CO ','HCHO ', &
'C2H6 ','C3H8 ','Me2CO ','MeCHO ', &
'C5H8 ','NO_aircrft'/)
- n_chem_tracers = 71 ! No chem tracers
+ n_chem_tracers = 73 ! No chem tracers
nr_therm = 220 ! thermal reactions
nr_phot = 55 ! photolytic (ATA)
@@ -325,7 +325,7 @@
'SO2_high ','NH3 ','DMS ','SO2_nat ', &
'BC_biomass','OC_biomass','NO_aircrft'/)
n_aero_tracers = 12
- n_chem_tracers = 71 ! No chem tracers
+ n_chem_tracers = 73 ! No chem tracers
IF (L_ukca_trophet) THEN
nr_therm = 241 ! thermal reactions
ELSE
Index: src/atmosphere/UKCA/ukca_set_nmspec.F90
===================================================================
--- src/atmosphere/UKCA/ukca_set_nmspec.F90 (revision 30974)
+++ src/atmosphere/UKCA/ukca_set_nmspec.F90 (revision 30979)
@@ -73,7 +73,7 @@
'BrO ','BrCl ','BrONO2 ','N2O ','HCl ', & !50
'HOCl ','HBr ','HOBr ','ClONO2 ','CFCl3 ', &
'CF2Cl2 ','MeBr ','N ','O(3P) ','MACRO2 ', & !60
-'MeCl ','CF2ClBr ','CCl4 ','CF2ClCFCl2','CHF2Cl ', &
+'MeCl ','CF2ClBr ','CCl4 ','ALICE ','BOB ', &
'MeCCl3 ','CF3Br ','H2OS ','CH2Br2 ','H2 ', & !70
'DMS ','SO2 ','H2SO4 ','MSA ','DMSO ', &
'NH3 ','CS2 ','COS ','H2S ','H ', & !80
Index: src/atmosphere/UKCA/ukca_constants.F90
===================================================================
--- src/atmosphere/UKCA/ukca_constants.F90 (revision 30974)
+++ src/atmosphere/UKCA/ukca_constants.F90 (revision 30979)
@@ -272,6 +272,10 @@
REAL, PARAMETER :: c_isosvoc2 = 2.3473 ! as C5H8
REAL, PARAMETER :: c_isosoa = 4.4874 ! 130.0
+! UKCA Tutorial tracers
+REAL, PARAMETER :: C_ALICE = 1.0000
+REAL, PARAMETER :: C_BOB = 1.0000
+
! molecular masses in g/mol of emitted species,
! for budget calculations
REAL, PARAMETER :: m_ho2 = 33.007
Index: src/atmosphere/UKCA/ukca_chem_strattrop.F90
===================================================================
--- src/atmosphere/UKCA/ukca_chem_strattrop.F90 (revision 30974)
+++ src/atmosphere/UKCA/ukca_chem_strattrop.F90 (revision 30979)
@@ -63,7 +63,7 @@
! ATA NLA CheST Chemistry v1.2
-TYPE(chch_t), PUBLIC :: chch_defs_strattrop_chem(1:75)=(/ &
+TYPE(chch_t), PUBLIC :: chch_defs_strattrop_chem(1:77)=(/ &
! 1
chch_t( 1,'O(3P) ', 1,'TR ','Ox ', 0, 0, 0), &
! 2
@@ -213,10 +213,14 @@
! 74
chch_t( 74,'O2 ', 1,'CT ',' ', 0, 0, 0), &
! 75
-chch_t( 75,'N2 ', 1,'CT ',' ', 0, 0, 0) &
+chch_t( 75,'N2 ', 1,'CT ',' ', 0, 0, 0), &
+! 76
+chch_t( 76,'ALICE ', 1,'TR ',' ', 0, 0, 0), &
+! 77
+chch_t( 77,'BOB ', 1,'TR ',' ', 0, 0, 0) &
/)
-TYPE(chch_t), PUBLIC :: chch_defs_strattrop_aer(1:87)=(/ &
+TYPE(chch_t), PUBLIC :: chch_defs_strattrop_aer(1:89)=(/ &
! 1
chch_t( 1,'O(3P) ', 1,'TR ','Ox ', 0, 0, 0), &
! 2
@@ -367,30 +371,34 @@
chch_t( 74,'O2 ', 1,'CT ',' ', 0, 0, 0), &
! 75
chch_t( 75,'N2 ', 1,'CT ',' ', 0, 0, 0), &
-! 76
-chch_t( 76,'DMS ', 1,'TR ',' ', 0, 0, 0), &
-! 77 DD:37,WD:31,EM:10
-chch_t( 77,'SO2 ', 1,'TR ',' ', 1, 1, 1), &
+! 76
+chch_t( 76,'ALICE ', 1,'TR ',' ', 0, 0, 0), &
+! 77
+chch_t( 77,'BOB ', 1,'TR ',' ', 0, 0, 0), &
! 78
-chch_t( 78,'H2SO4 ', 1,'TR ',' ', 0, 0, 0), &
-! 79
-chch_t( 79,'MSA ', 1,'TR ',' ', 0, 0, 0), &
-! 80 DD:38,WD:32
-chch_t( 80,'DMSO ', 1,'TR ',' ', 1, 1, 0), &
-! 81 DD:39,WD:33,EM:11
-chch_t( 81,'NH3 ', 1,'TR ',' ', 1, 1, 1), &
-! 82
-chch_t( 82,'CS2 ', 1,'TR ',' ', 0, 0, 0), &
-! 83
-chch_t( 83,'COS ', 1,'TR ',' ', 0, 0, 0), &
+chch_t( 78,'DMS ', 1,'TR ',' ', 0, 0, 0), &
+! 79 DD:37,WD:31,EM:10
+chch_t( 79,'SO2 ', 1,'TR ',' ', 1, 1, 1), &
+! 80
+chch_t( 80,'H2SO4 ', 1,'TR ',' ', 0, 0, 0), &
+! 81
+chch_t( 81,'MSA ', 1,'TR ',' ', 0, 0, 0), &
+! 82 DD:38,WD:32
+chch_t( 82,'DMSO ', 1,'TR ',' ', 1, 1, 0), &
+! 83 DD:39,WD:33,EM:11
+chch_t( 83,'NH3 ', 1,'TR ',' ', 1, 1, 1), &
! 84
-chch_t( 84,'H2S ', 1,'TR ',' ', 0, 0, 0), &
+chch_t( 84,'CS2 ', 1,'TR ',' ', 0, 0, 0), &
! 85
-chch_t( 85,'SO3 ', 1,'TR ',' ', 0, 0, 0), &
-! 86 DD:40, EM: 12
-chch_t( 86,'Monoterp ', 1,'TR ',' ', 1, 0, 1), &
-! 87 DD:41,WD:34
-chch_t( 87,'Sec_Org ', 1,'TR ',' ', 1, 1, 0) &
+chch_t( 85,'COS ', 1,'TR ',' ', 0, 0, 0), &
+! 86
+chch_t( 86,'H2S ', 1,'TR ',' ', 0, 0, 0), &
+! 87
+chch_t( 87,'SO3 ', 1,'TR ',' ', 0, 0, 0), &
+! 88 DD:40, EM: 12
+chch_t( 88,'Monoterp ', 1,'TR ',' ', 1, 0, 1), &
+! 89 DD:41,WD:34
+chch_t( 89,'Sec_Org ', 1,'TR ',' ', 1, 1, 0) &
/)
TYPE(ratb_t) :: ratb_defs_strattrop_chem(198)
Index: src/atmosphere/UKCA/ukca_setup_chem_mod.F90
===================================================================
--- src/atmosphere/UKCA/ukca_setup_chem_mod.F90 (revision 30974)
+++ src/atmosphere/UKCA/ukca_setup_chem_mod.F90 (revision 30979)
@@ -248,8 +248,8 @@
l_ukca_strattrop = .TRUE.
l_ukca_advh2o = .TRUE.
ukca_int_method = int_method_nr
- jpctr = 71
- jpspec = 75
+ jpctr = 73
+ jpspec = 77
jpbk = 198
jptk = 24
jppj = 56
Index: src/atmosphere/UKCA/ukca_cspecies.F90
===================================================================
--- src/atmosphere/UKCA/ukca_cspecies.F90 (revision 30974)
+++ src/atmosphere/UKCA/ukca_cspecies.F90 (revision 30979)
@@ -278,6 +278,9 @@
WHERE (advt == 'ORGNIT ') c_species = c_orgnit
WHERE (advt == 'PASSIVE O3') c_species = 1.0
WHERE (advt == 'AGE OF AIR') c_species = 1.0
+! UKCA Tutorial Tracers
+WHERE (advt == 'ALICE ') c_species = C_ALICE
+WHERE (advt == 'BOB ') c_species = C_BOB
! non-advected tracers
c_na_species=0.0
Index: rose-meta/um-atmos/HEAD/etc/stash/STASHmaster/STASHmaster-meta.conf
===================================================================
--- rose-meta/um-atmos/HEAD/etc/stash/STASHmaster/STASHmaster-meta.conf (revision 30974)
+++ rose-meta/um-atmos/HEAD/etc/stash/STASHmaster/STASHmaster-meta.conf (revision 30979)
@@ -13144,6 +13144,14 @@
description=O3P MASS MIXING RATIO AFTER TIMESTEP
help=Oxygen atom (ground state) Mass Mixing Ratio in kg/kg(Air)
+[stashmaster:code(34064)]
+description=ALICE MASS MIXING RATIO AFTER TIMESTEP
+help=UKCA Tutiorial Tracer ALICE Mass Mixing Ratio in kg/kg(Air)
+
+[stashmaster:code(34065)]
+description=BOB MASS MIXING RATIO AFTER TIMESTEP
+help=UKCA Tutiorial Tracer BOB Mass Mixing Ratio in kg/kg(Air)
+
[stashmaster:code(34070)]
description=H2 MASS MIXING RATIO AFTER TIMESTEP
help=Hydrogen Mass Mixing Ratio in kg/kg(Air)
@@ -18069,6 +18077,16 @@
description=O3P MASS MIXING RATIO ON PRESS LEVS
help=Oxygen atom (ground state) Mass Mixing Ratio in kg/kg(Air)
+[stashmaster:code(51064)]
+description=ALICE MASS MIXING RATIO AFTER TIMESTEP
+help=UKCA Tutiorial Tracer ALICE Mass Mixing Ratio in kg/kg(Air)
+ =on pressure levels
+
+[stashmaster:code(51065)]
+description=BOB MASS MIXING RATIO AFTER TIMESTEP
+help=UKCA Tutiorial Tracer BOB Mass Mixing Ratio in kg/kg(Air)
+ =on pressure levels
+
[stashmaster:code(51070)]
description=H2 MASS MIXING RATIO ON PRESS LEVS
help=Hydrogen Mass Mixing Ratio in kg/kg(Air)
Index: rose-meta/um-atmos/HEAD/etc/stash/STASHmaster/STASHmaster_A
===================================================================
--- rose-meta/um-atmos/HEAD/etc/stash/STASHmaster/STASHmaster_A (revision 30974)
+++ rose-meta/um-atmos/HEAD/etc/stash/STASHmaster/STASHmaster_A (revision 30979)
@@ -13933,6 +13933,18 @@
4| 1 | 0 | -99 -99 -99 -99 -99 -99 -99 -99 -99 -99 |
5| 0 | 2160 | 0 | 65 | 0 | 0 | 0 | 0 | 0 |
#
+1| 1 | 34 | 64 |ALICE MASS MIXING RATIO AFTER TS |
+2| 2 | 0 | 1 | 1 | 2 | 40 | 11 | 0 | 0 | 0 | 0 |
+3| 000000000000000000000000010000 | 00000000000000000001 | 1 |
+4| 1 | 0 | -99 -99 -99 -99 -99 -99 -99 -99 -99 -99 |
+5| 0 | 2164 | 0 | 65 | 0 | 0 | 0 | 0 | 0 |
+#
+1| 1 | 34 | 65 |BOB MASS MIXING RATIO AFTER TS |
+2| 2 | 0 | 1 | 1 | 2 | 40 | 11 | 0 | 0 | 0 | 0 |
+3| 000000000000000000000000010000 | 00000000000000000001 | 1 |
+4| 1 | 0 | -99 -99 -99 -99 -99 -99 -99 -99 -99 -99 |
+5| 0 | 2165 | 0 | 65 | 0 | 0 | 0 | 0 | 0 |
+#
1| 1 | 34 | 69 |CH3OH MASS MIXING RATIO AFTER TS |
2| 2 | 0 | 1 | 1 | 2 | 40 | 11 | 0 | 0 | 0 | 0 |
3| 000000000000000000000000000100 | 00000000000000000001 | 1 |
@@ -21785,6 +21797,18 @@
4| 1 | 0 | -99 -99 -99 -99 -99 -99 -99 -99 -99 -99 |
5| 0 | 2160 | 0 | 8 | 0 | 0 | 0 | 0 | 0 |
#
+1| 1 | 51 | 64 |ALICE MASS MIX RATIO ON PRESS LEVS |
+2| 0 | 0 | 1 | 1 | 3 | 1 | 2 | 0 | 0 | 0 | 1 |
+3| 000000000000000000000000010000 | 00000000000000000001 | 3 |
+4| 1 | 0 | -99 -99 -99 -99 -99 -99 -99 -99 -99 -99 |
+5| 0 | 2164 | 0 | 8 | 0 | 0 | 0 | 0 | 0 |
+#
+1| 1 | 51 | 65 |BOB MASS MIX RATIO ON PRESS LEVS |
+2| 0 | 0 | 1 | 1 | 3 | 1 | 2 | 0 | 0 | 0 | 1 |
+3| 000000000000000000000000010000 | 00000000000000000001 | 3 |
+4| 1 | 0 | -99 -99 -99 -99 -99 -99 -99 -99 -99 -99 |
+5| 0 | 2165 | 0 | 8 | 0 | 0 | 0 | 0 | 0 |
+#
1| 1 | 51 | 69 |CH3OH MASS MIX RATIO ON PRESS LEVS |
2| 0 | 0 | 1 | 1 | 3 | 1 | 2 | 0 | 0 | 0 | 1 |
3| 000000000000000000000000000100 | 00000000000000000001 | 3 |
These differences can be found in the file /home/ukca/Tutorial/worked_solutions/Task4.1/task4.1.rose.diff on PUMA.
If you open the .pk file in Xconv, you should see the following fields:
0 : 192 144 36 1 O3 MASS MIXING RATIO 1 : 192 144 36 1 Molar flux density 2 : 192 144 85 1 O3 MASS MIXING RATIO 3 : 192 144 85 1 Field code = 2164 4 : 192 144 85 1 Field code = 2165
Sample output from this task can be found at /work/n02/n02/ukca/Tutorial/vn10.4/sample_output/Task4.1/ai084a.pk19880901 on ARCHER.
Written by Luke Abraham 2016