Difference between revisions of "UKCA Chemistry and Aerosol Tutorials: Things to know before you start the vn10.9 practicals"
| (11 intermediate revisions by the same user not shown) | |||
| Line 19: | Line 19: | ||
https://code.metoffice.gov.uk/doc/um/latest/um-training/index.html |
https://code.metoffice.gov.uk/doc/um/latest/um-training/index.html |
||
| − | Also, if you do not have a PUMA or ARCHER account, you can use the Met Office provided Virtual Machine to run the VM |
+ | Also, if you do not have a PUMA or ARCHER account, you can use the Met Office provided Virtual Machine to run the VM, provided here: |
| + | https://github.com/metomi/metomi-vms |
||
| ⚫ | |||
| + | |||
| + | You should follow the instructions in [https://code.metoffice.gov.uk/doc/um/latest/umdp.html UMDP X10]. |
||
| + | |||
| ⚫ | |||
===Accounts=== |
===Accounts=== |
||
| Line 55: | Line 59: | ||
!style="text-align:top;"|Comments |
!style="text-align:top;"|Comments |
||
|- |
|- |
||
| − | | ARCHER || vn10.9 N48L38 Intel Compiler || '''u-as101''' || lukeabraham || May be [[#Manual_Compiling|compiled manually (recommended)]] |
+ | | ARCHER || vn10.9 N48L38 Intel Compiler || '''u-as101''' || lukeabraham || May be [[#Manual_Compiling (ARCHER)|compiled manually (recommended)]] |
|- |
|- |
||
| vm || vn10.9 N48L38 GNU Compiler || '''u-as159''' || lukeabraham || 1x2 decomposition |
| vm || vn10.9 N48L38 GNU Compiler || '''u-as159''' || lukeabraham || 1x2 decomposition |
||
| + | |- |
||
| + | | XC40 (Monsoon2/NEXCS) || vn10.9 N48L38 GNU Compiler || '''u-au234''' || lukeabraham || 4x9 decomposition (36-core) |
||
|- |
|- |
||
|} |
|} |
||
| Line 65: | Line 71: | ||
To set-up using the UM with Rose & Cylc correctly, go to |
To set-up using the UM with Rose & Cylc correctly, go to |
||
| − | * |
+ | * https://ncas-cms.github.io/um-training/getting-setup.html |
and follow the instructions from '''Section 1.2''' onwards if you are using Linux/Mac (after opening a terminal). |
and follow the instructions from '''Section 1.2''' onwards if you are using Linux/Mac (after opening a terminal). |
||
| Line 79: | Line 85: | ||
When you have copied the suite, you will need to remember to change the username & tic code in ''suite conf <math>\rightarrow</math> Machine Options''. |
When you have copied the suite, you will need to remember to change the username & tic code in ''suite conf <math>\rightarrow</math> Machine Options''. |
||
| + | ===Additional Requirements (Virtual Machine)=== |
||
| ⚫ | |||
| + | |||
| + | ====Tutorial Data==== |
||
| + | |||
| + | To be able to do Tutorials 5 and 10 you will need emissions data and python scripts. These files, as well as worked solutions for all the Rose and code changes can be obtained by running the following command from the <code>home/</code> directory of your VM: |
||
| + | |||
| + | wget -q --show-progress http://gws-access.ceda.ac.uk/public/ukca/UKCA_Tutorial_vn109.tgz |
||
| + | |||
| + | There is 880MB to download, so it might take some time. |
||
| + | |||
| + | Once the download has finished, you can extract the archive by running |
||
| + | |||
| + | tar -xzvf UKCA_Tutorial_vn109.tgz |
||
| + | |||
| + | This will make a directory called <code>Tutorial/</code> containing 1.4GB of files. If you <code>cd</code> into this you can check that the files have not been corrupted by running the following command: |
||
| + | |||
| + | md5sum --status -c vn109.md5 |
||
| + | |||
| + | ====Xconv==== |
||
| + | |||
| + | You will need to download [http://cms.ncas.ac.uk/documents/xconv/ Xconv] (<code>xconv1.93</code>) from here: |
||
| + | |||
| + | * http://cms.ncas.ac.uk/documents/xconv/_downloads/xconv1.93_linux_x86_64.tar.gz |
||
| + | |||
| + | You can download this by |
||
| + | |||
| + | wget <nowiki>http://cms.ncas.ac.uk/documents/xconv/_downloads/xconv1.93_linux_x86_64.tar.gz</nowiki> |
||
| + | |||
| + | Download it to your $HOME/bin on the VM, cd into this directory, <code>tar -zxvf</code> the tar-ball, and then |
||
| + | |||
| + | ln -s xconv1.93 xconv |
||
| + | ln -s xconv1.93 convsh |
||
| + | |||
| + | ====Iris==== |
||
| + | |||
| + | There is an '''<code>install-iris</code>''' script provided, but you will need to set-up modules yourself to be able to use it properly. The anaconda install breaks Rose if put in your PATH, however, there is now an alias |
||
| + | |||
| + | '''conda''' |
||
| + | |||
| + | which will open a new terminal with all the anaconda python packages in its $PATH. This will allow you to use Rose in one terminal and Iris in another. |
||
| + | |||
| + | I have found a handy way to use python is to use [https://ipython.org/ <code>ipython</code>] with the following arguments |
||
| + | |||
| + | ipython --pylab --logfile=ipython-`date +\"%Y%m%d-%H%M%S\"`.py" |
||
| + | |||
| + | I have aliased this to |
||
| + | |||
| + | pylab |
||
| + | |||
| + | in my <code>.bashrc</code>. The |
||
| + | |||
| + | --pylab |
||
| + | |||
| + | sets-up a MatLab-type environment (numpy, scipy, matplotlib all loaded using standard shortcuts), and |
||
| + | |||
| + | --logfile=ipython-`date +\"%Y%m%d-%H%M%S\"`.py |
||
| + | |||
| + | means that all commands are saved to a file of the format |
||
| + | |||
| + | ipython-YYMMDD-HHMMSS.py |
||
| + | |||
| ⚫ | |||
<gallery widths=300px heights=200px> |
<gallery widths=300px heights=200px> |
||
| Line 107: | Line 174: | ||
==Scripts== |
==Scripts== |
||
| − | There are a number of scripts provided for these tutorials |
+ | There are a number of scripts provided for these tutorials. These can be found on GitHub here: |
* https://github.com/theabro/ukca |
* https://github.com/theabro/ukca |
||
| + | |||
| + | They are also available on ARCHER. |
||
[[UKCA Chemistry and Aerosol Tutorials at vn10.9]] |
[[UKCA Chemistry and Aerosol Tutorials at vn10.9]] |
||
Latest revision as of 09:08, 20 May 2020
UKCA Chemistry and Aerosol Tutorials at vn10.9
You will need to have the details of your PUMA and ARCHER accounts to hand.
Before you begin the UKCA Tutorials
These tutorials assume that you are familiar with FCM, and training on this can be found here:
http://cms.ncas.ac.uk/wiki/Fcm
The UM FCM tutorial that is linked from this page uses the old user interface (the "UMUI"), so will be less relevant here.
You should also familiarise yourself with Rose & Cylc, and more information on these can be found here:
http://cms.ncas.ac.uk/wiki/RoseCylc
The Met Office provide some on-line UM specific Rose training here:
https://code.metoffice.gov.uk/doc/um/latest/um-training/index.html
Also, if you do not have a PUMA or ARCHER account, you can use the Met Office provided Virtual Machine to run the VM, provided here:
https://github.com/metomi/metomi-vms
You should follow the instructions in UMDP X10.
It is possible to run a simple UKCA suite in the VM, but note that this requires at least 6GB of RAM to run successfully.
Accounts
To run these tutorials the minimum you will need is
- an account on the Met Office Science Repository Service
which will then allow you to run the tutorials in a Virtual Machine.
To run these on ARCHER you will need
- a PUMA account
- an ARCHER account
For information on how to get these, please see the Getting Started with UKCA page.
If you are attending the face-to-face training workshop you will be provided with ARCHER training accounts, although you still need your own PUMA account.
Developing code for the UM
During this tutorial you will be creating tickets on the Met Office Science Repository Service, and making code changes to UM branches. Please note that this is a live system, used by scientists all over the world to develop their code. However, please don't worry about breaking anything, as there are practices in place to prevent this.
For more information about developing for the UM, please see the Working Practices for UM Development with Rose, FCM and trac.
UKCA Training Suite
| Machine | UM Version/Configuration | Suite ID | Owner | Comments |
|---|---|---|---|---|
| ARCHER | vn10.9 N48L38 Intel Compiler | u-as101 | lukeabraham | May be compiled manually (recommended) |
| vm | vn10.9 N48L38 GNU Compiler | u-as159 | lukeabraham | 1x2 decomposition |
| XC40 (Monsoon2/NEXCS) | vn10.9 N48L38 GNU Compiler | u-au234 | lukeabraham | 4x9 decomposition (36-core) |
These suites are based on the um_n48_ukca_eg app, which is available in the MetUM trunk (in rose-stem/app/). A ticket giving more details on these (and the code changes) can be found at: um:#3578.
To set-up using the UM with Rose & Cylc correctly, go to
and follow the instructions from Section 1.2 onwards if you are using Linux/Mac (after opening a terminal).
Note: when entering MOSRS password, a rose window will open behind everything all your other windows. You will also need to enter your username here to complete the process.
When you have completed this, you should launch the Rose browser by typing
rosie go &
and there you should search for training suite and make a copy of it.
When you have copied the suite, you will need to remember to change the username & tic code in suite conf Machine Options.
Additional Requirements (Virtual Machine)
Tutorial Data
To be able to do Tutorials 5 and 10 you will need emissions data and python scripts. These files, as well as worked solutions for all the Rose and code changes can be obtained by running the following command from the home/ directory of your VM:
wget -q --show-progress http://gws-access.ceda.ac.uk/public/ukca/UKCA_Tutorial_vn109.tgz
There is 880MB to download, so it might take some time.
Once the download has finished, you can extract the archive by running
tar -xzvf UKCA_Tutorial_vn109.tgz
This will make a directory called Tutorial/ containing 1.4GB of files. If you cd into this you can check that the files have not been corrupted by running the following command:
md5sum --status -c vn109.md5
Xconv
You will need to download Xconv (xconv1.93) from here:
You can download this by
wget http://cms.ncas.ac.uk/documents/xconv/_downloads/xconv1.93_linux_x86_64.tar.gz
Download it to your $HOME/bin on the VM, cd into this directory, tar -zxvf the tar-ball, and then
ln -s xconv1.93 xconv ln -s xconv1.93 convsh
Iris
There is an install-iris script provided, but you will need to set-up modules yourself to be able to use it properly. The anaconda install breaks Rose if put in your PATH, however, there is now an alias
conda
which will open a new terminal with all the anaconda python packages in its $PATH. This will allow you to use Rose in one terminal and Iris in another.
I have found a handy way to use python is to use ipython with the following arguments
ipython --pylab --logfile=ipython-`date +\"%Y%m%d-%H%M%S\"`.py"
I have aliased this to
pylab
in my .bashrc. The
--pylab
sets-up a MatLab-type environment (numpy, scipy, matplotlib all loaded using standard shortcuts), and
--logfile=ipython-`date +\"%Y%m%d-%H%M%S\"`.py
means that all commands are saved to a file of the format
ipython-YYMMDD-HHMMSS.py
Manual Compiling (ARCHER)
The UKCA Training Suite has been altered slightly to allow it to be easily manually compiled. This can be selected as an option in suite conf UM hosts. With this set to true, when the suite runs the fcm_make2 task will enter the submit-failed state. Then, on ARCHER, you should then change directory into
$HOME/cylc-run/[SUITE-ID]/log/job/1/fcm_make2/NN
and compile the um-atmos and um-recon executables by typing
./job
This will take around 7-12 minutes to compile. When it has finished successfully you will get the following message in the terminal, just before the prompt:
JOB SCRIPT EXITING (TASK SUCCEEDED)
You can follow the progress of this process in detail by tail-ing the following file
$HOME/cylc-run/[SUITE-ID]/share/fcm_make/fcm-make2.log
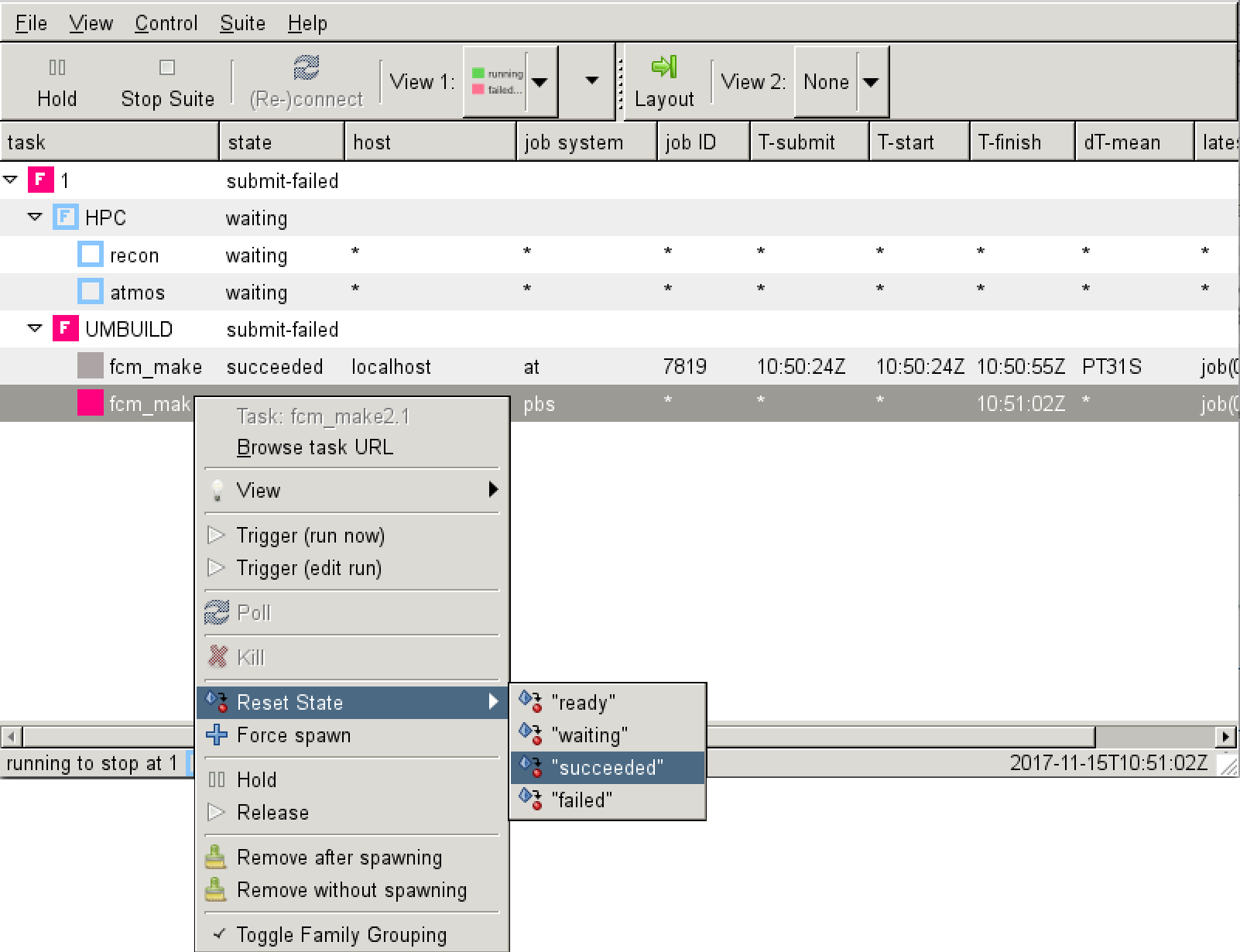
Once this step has completed, you can send the rest of the suite off by going back to the Gcylc window and right-clicking on the fcm_make2 task and click Reset State "succeeded". This will then allow the rest of the tasks to start in sequence.
Note: manually compiling in this way means that you will no longer have the job.out or job.err files. You should instead look in the fcm-make2.log file.
Scripts
There are a number of scripts provided for these tutorials. These can be found on GitHub here:
They are also available on ARCHER.
UKCA Chemistry and Aerosol Tutorials at vn10.9
Written by Luke Abraham 2017.