UKCA Chemistry and Aerosol vn10.9 Tutorial 9
UKCA Chemistry and Aerosol Tutorials at vn10.9
What you will learn in this Tutorial
In this tutorial you will learn about the UKCA diagnostics package and the different diagnostics that you can output using it. You will also learn how to add new diagnostics from the new reactions and deposition that you have added.
Task 9.1: Output new diagnostics
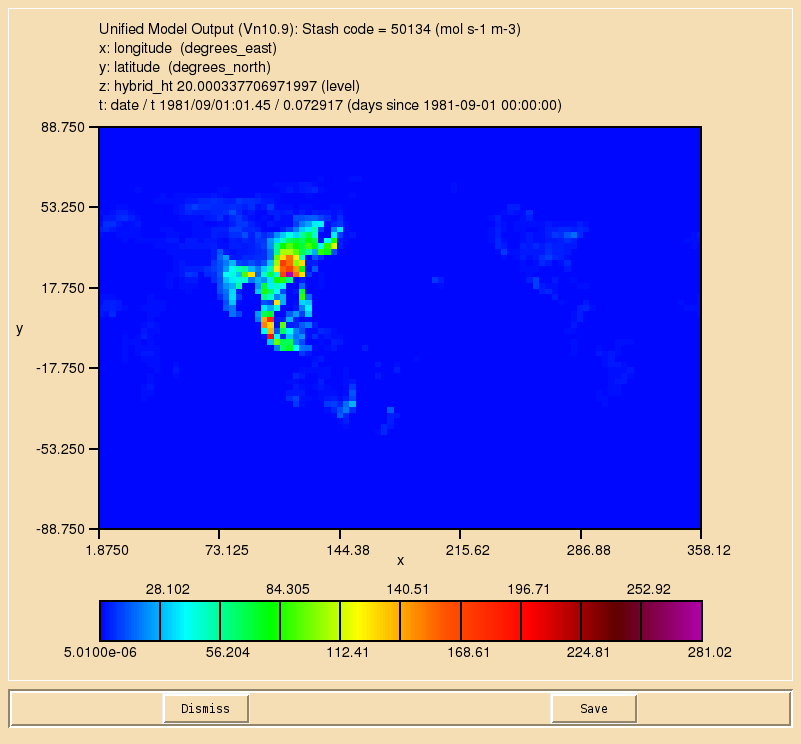
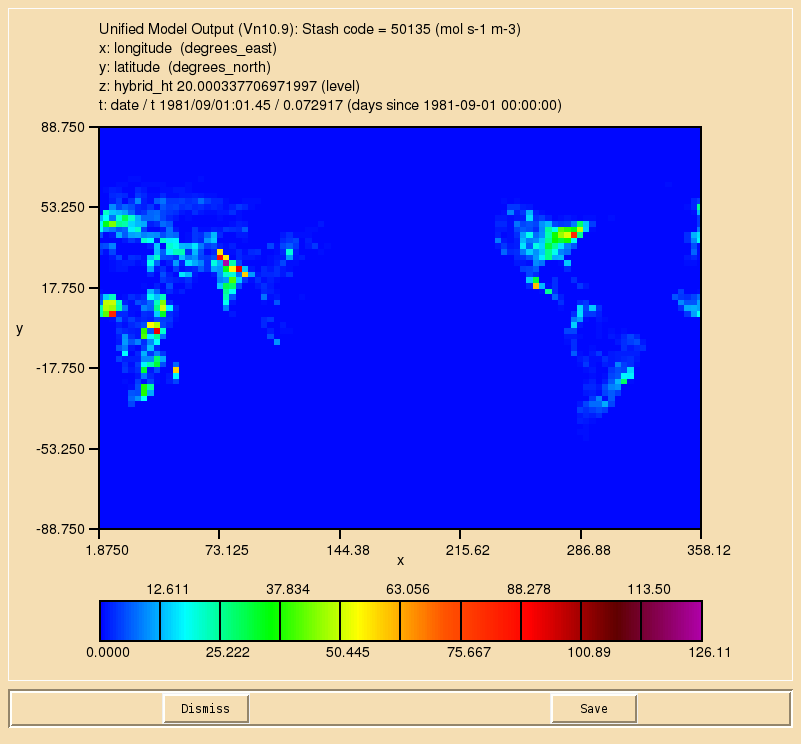
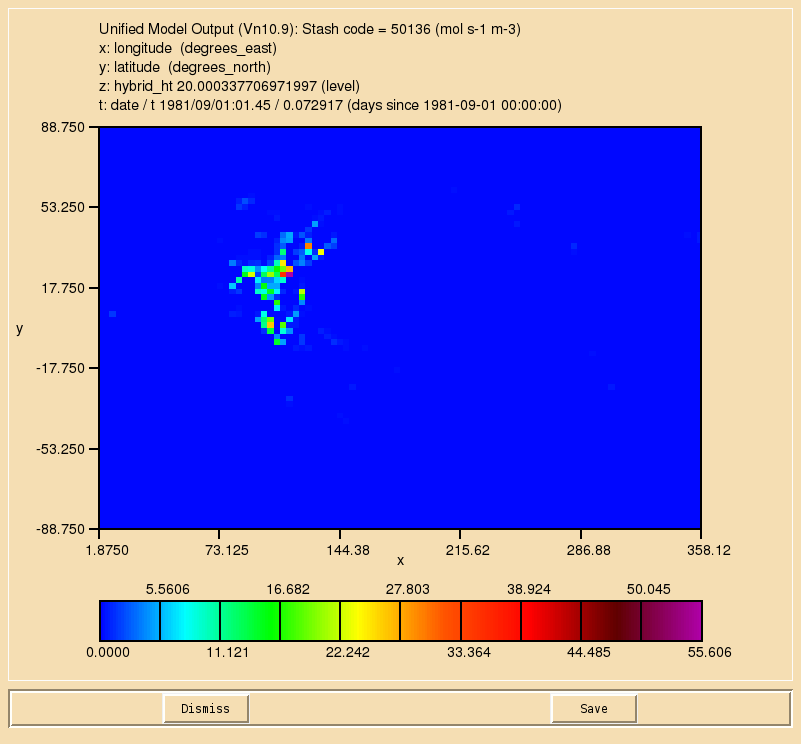
TASK 9.1: Output diagnostics of the reaction to STASH code 50134, the dry deposition of ALICE to STASH code 50135, and the wet deposition of BOB to 50136. They should be outputted as a 3-hour mean to the pa/UPA stream.
Adding New UKCA Diagnostics
If you are using one of the chemistry schemes that uses ASAD (e.g. CheT/TropIsop, CheS/Strat, CheST/StratTrop) then you can make use of the ASAD Reaction Flux Diagnostics module (held in asad_chem_flux_diags.F90). These allow you to straight-forwardly output new reaction and deposition fluxes.
To output new diagnostics you will first need to define them in the asad_flux_dat.F90 module, and then create new STASHmaster file specifications for them.
During this tutorial you will be tasked with outputting the reaction and deposition fluxes that you have added in to your branch.
Flux Definitions in asad_flux_dat.F90
Within the asad_flux_dat.F90 module the diagnostics are defined in blocks with the format
asad_flux_defn('Diagnostic type',STASH code,'Diagnostic specification',Mask,Reaction number,Number of species, & (/'Species/Reactant 1','Reactant 2'/), & (/'Product 1','Product 2','Product 3','Product 4'/)), &
Which have the following meaning:
Diagnostic Type
This is a three character string which defines what type of diagnostic is being requested. This can take the values
- RXN to output the flux through a reaction (in moles/gridcell/s)
- DEP to output the deposition flux of a species (in moles/gridcell/s)
- NET to output the net chemical tendency of a species (in moles/gridcell/s)
- STE to output the net dynamical tendency of a species (in moles/gridcell/s)
- MAS to output the mass of the atmosphere (in kg/gridcell)
- PSC to output polar stratospheric cloud diagnostics (1 when the gridcell contains a PSC, 0 otherwise - monthly mean field will be a fraction in range 0 → 1)
- TPM to output the tropospheric mask (1 for troposphere, 0 otherwise - monthly mean field will be a fraction in range 0 → 1)
- OUT to output a tracer in mmr. Only really useful if the field is masked to give the tropospheric concentration only (see the discussion of the Mask option)
STASH Code
This is a 5 digit integer defining the STASH code that the diagnostic will be outputted to (e.g. 50001). Currently this must be in section 50.
Diagnostic Specification
This is a one character string which is needed to further define what diagnostic is required. If it isn't needed then it should just be set to X or left blank.
- RXN
- B to output the flux through a bimolecular reaction
- T to output the flux through a termolecular reaction
- H to output the flux through a heterogeneous reaction
- J to output the flux through a photolysis reaction
- DEP
- D to output the dry deposition flux
- W to output the wet deposition flux
- PSC
- 1 to output the fraction of Type 1 PSCs
- 2 to output the fraction of Type 2 PSCs
Mask
This is a logical which defines whether only the tropospheric values of the diagnostic are outputted (.TRUE.) or not (.FALSE.). It is calculated every timestep.
For the STE diagnostic this is required if you wish to output the diagnosed stratosphere-troposphere exchange of a species. For the OUT diagnostic this can be used to output only the tropospheric concentration of a tracer. This is also used in the calculation of the of the TPM diagnostic.
Reaction number
This is an integer, and should only be used in the special case of there being two (or more) reactions with the exactly the same reactants and products, but with different rate coefficients. In this case the first reaction in the list would be given number 1 and the second 2 etc. If this is not needed then it should be set to 0 (which will be usual for most reactions).
Number of Species
This is an integer, and should give the total number of species, so this will be 1 for diagnostics such as DEP, STE, NET etc., which only consider a single species, and the total number of reactants and products for diagnostics RXN and RTE.
Species
This is a 10-character string giving the exact name of the species that the diagnostic should be considered for (including capitalisation). This is only used for the DEP, NET, STE, and OUT. For the RXN and RTE diagnostics the full list of reactants and products should be given (see below). For the MAS, PSC, and TPM diagnostics this isn't needed and could either be set to XXX or left blank. If it is needed the other reactant/product slots should be left blank.
Reactants and Products
These are 10-character strings, and should be as the reaction is defined in the ukca_chem_master.F90 module.
Summing Diagnostics
If you define more than one diagnostic to be output to the same STASH code, then the diagnostic routines will sum these diagnostics together. This can be useful (e.g., if you wanted to output the sum of all NO+RO2 reactions to one STASH item), but can be problematic if you accidentally output two fields to the same STASH code, as this will give strange results!
Changes to asad_flux_dat.F90
After you have defined your new diagnostics at the top of this module, you will need to make sure that they have been added correctly to the asad_chemical_fluxes array, which is defined in the asad_load_default_fluxes subroutine held in the asad_flux_dat.F90 module. You will need to increment the size of this array, given by the n_chemical_fluxes parameter.
Changes to asad_chem_flux_diags.F90 (and other UKCA routines)
You will need to edit asad_chem_flux_diags.F90 module if you want to output a new type of diagnostic. This can be quite involved, but you can look at existing routines to see how things are done. You will also need to add code into the main UKCA routines the pass the data through, e.g.
! 3D flux diagnostics
IF (L_asad_use_chem_diags .AND. &
((L_asad_use_flux_rxns .OR. L_asad_use_rxn_rates) .OR. &
(L_asad_use_wetdep .OR. L_asad_use_drydep))) &
CALL asad_chemical_diagnostics(row_length,rows, &
model_levels,ix,jy,k,secs_per_step,volume,ierr)
STASHmaster file
While the diagnostics are defined in asad_flux_dat.F90 they are turned on by requesting the item through STASH. To do this you will need to edit the STASHmaster_A file in your branch.
Most UKCA diagnostics are 3D, although some, such as emissions (which are outputted in a different way), are 2D. You should take care with the STASH settings in STASHmaster_A between these types, as there some differences that will need to be considered.
Rose Changes
If you have not done so, you will also need to make sure that you use your new STASHmaster_A file in Rose, as is explained in detail in Tutorial 4.
After you have made your STASmaster_A file changes, you will need to add these diagnostics into STASH, as per Tutorial 3. Remember to run the TidyStashTransform macro. As you will have included your branch's STASHmaster_A file using @HEAD, you won't need to make any further changes to Rose, but you will need to make sure that you have committed your UM branch prior to running. Because the file is taken from fcm:um.xm_br you will need to make sure that the revision has synced to the PUMA mirror (i.e. by waiting a few minutes).
Solution to Task 9.1: Output new diagnostics
You were given the task
- Output diagnostics of the reaction to STASH code 50134, the dry deposition of ALICE to STASH code 50135, and the wet deposition of BOB to 50136. They should be outputted as a 3-hour mean to the pa/UPA stream.
For a working Rose suite that has completed this task, please see
- ARCHER:
u-as292@60289 - vm:
u-as297@60286
The specific Rose changes made are:
- ARCHER: https://code.metoffice.gov.uk/trac/roses-u/changeset/60289/a/s/2/9/2/trunk
- vm: https://code.metoffice.gov.uk/trac/roses-u/changeset/60286/a/s/2/9/7/trunk
The specific Rose changes made are:
ARCHER:
Index: app/um/rose-app.conf =================================================================== --- app/um/rose-app.conf (revision 60205) +++ app/um/rose-app.conf (revision 60289) @@ -3425,6 +3425,30 @@ tim_name='T3HMN' use_name='UPA' +[namelist:umstash_streq(50134_de3d001f)] +dom_name='DALLTH' +isec=50 +item=134 +package= +tim_name='T3HMN' +use_name='UPA' + +[namelist:umstash_streq(50135_8528080c)] +dom_name='DALLTH' +isec=50 +item=135 +package= +tim_name='T3HMN' +use_name='UPA' + +[namelist:umstash_streq(50136_fb2dacd3)] +dom_name='DALLTH' +isec=50 +item=136 +package= +tim_name='T3HMN' +use_name='UPA' + [namelist:umstash_streq(50156_4ce4e1d4)] dom_name='DIAG' isec=50 Index: app/fcm_make/rose-app.conf =================================================================== --- app/fcm_make/rose-app.conf (revision 60205) +++ app/fcm_make/rose-app.conf (revision 60289) @@ -42,4 +42,4 @@ stash_version=1A timer_version=3A um_rev=vn10.9 -um_sources=branches/dev/lukeabraham/vn10.9_UKCA_Tutorial_Solns@46696 +um_sources=branches/dev/lukeabraham/vn10.9_UKCA_Tutorial_Solns@46718
These differences can be found in the file /home/ukca/Tutorial/vn10.9/worked_solutions/Task9.1/Task9.1_rose.patch on PUMA.
vm:
Index: app/um/rose-app.conf =================================================================== --- app/um/rose-app.conf (revision 60206) +++ app/um/rose-app.conf (revision 60286) @@ -4017,6 +4017,30 @@ tim_name='T3HMN' use_name='UPA' +[namelist:umstash_streq(50134_de3d001f)] +dom_name='DALLTH' +isec=50 +item=134 +package= +tim_name='T3HMN' +use_name='UPA' + +[namelist:umstash_streq(50135_8528080c)] +dom_name='DALLTH' +isec=50 +item=135 +package= +tim_name='T3HMN' +use_name='UPA' + +[namelist:umstash_streq(50136_fb2dacd3)] +dom_name='DALLTH' +isec=50 +item=136 +package= +tim_name='T3HMN' +use_name='UPA' + [namelist:umstash_streq(50156_4ce4e1d4)] dom_name='DIAG' isec=50 Index: app/fcm_make/rose-app.conf =================================================================== --- app/fcm_make/rose-app.conf (revision 60206) +++ app/fcm_make/rose-app.conf (revision 60286) @@ -42,4 +42,4 @@ stash_version=1A timer_version=3A um_rev=vn10.9 -um_sources=branches/dev/lukeabraham/vn10.9_UKCA_Tutorial_Solns@46696 +um_sources=branches/dev/lukeabraham/vn10.9_UKCA_Tutorial_Solns@46718
The specific UM changes made are:
Index: src/atmosphere/UKCA/asad_flux_dat.F90
===================================================================
--- src/atmosphere/UKCA/asad_flux_dat.F90 (revision 46696)
+++ src/atmosphere/UKCA/asad_flux_dat.F90 (revision 46718)
@@ -97,7 +97,7 @@
CHARACTER(LEN=10) :: blank0 = ' ' ! Defines null product
! Number of chemical fluxes defined below
-INTEGER, PARAMETER :: n_chemical_fluxes = 310
+INTEGER, PARAMETER :: n_chemical_fluxes = 313
TYPE(asad_flux_defn), ALLOCATABLE, SAVE :: asad_chemical_fluxes(:)
@@ -1285,6 +1285,19 @@
(/' ',' ',' ',' '/)) &
/)
+TYPE(asad_flux_defn), PARAMETER, PUBLIC :: &
+ ukca_tutorial_fluxes(3) = (/ &
+asad_flux_defn('RXN',50134,'B',.FALSE.,0,4, &
+(/'ALICE ','OH '/), &
+(/'BOB ','Sec_Org ',' ',' '/)), &
+asad_flux_defn('DEP',50135,'D',.FALSE.,0,1, &
+(/'ALICE ',' '/), &
+(/' ',' ',' ',' '/)), &
+asad_flux_defn('DEP',50136,'W',.FALSE.,0,1, &
+(/'BOB ',' '/), &
+(/' ',' ',' ',' '/)) &
+/)
+
TYPE(asad_flux_defn), PUBLIC :: asad_aerosol_chem(16)
PUBLIC :: asad_load_default_fluxes
@@ -1367,7 +1380,8 @@
asad_ro2ro2_reacn, & ! 8 263
asad_ch4_oxidn, & ! 6 269
asad_o1d_prod, & ! 3 272
- asad_h2o_budget & ! 38 307
+ asad_h2o_budget, & ! 38 307
+ ukca_tutorial_fluxes & ! 3
/)
IF (printstatus > PrStatus_Normal) THEN
Index: rose-meta/um-atmos/HEAD/etc/stash/STASHmaster/STASHmaster-meta.conf
===================================================================
--- rose-meta/um-atmos/HEAD/etc/stash/STASHmaster/STASHmaster-meta.conf (revision 46696)
+++ rose-meta/um-atmos/HEAD/etc/stash/STASHmaster/STASHmaster-meta.conf (revision 46718)
@@ -21102,6 +21102,21 @@
=number of moles of Ox being produced by this reaction per second in the
=whole model.
+[stashmaster:code(50134)]
+description=RXN FLUX: ALICE+OH->BOB+Sec_Org
+help=Flux through ALICE+OH->BOB+Sec_Org reaction
+ =moles/s
+
+[stashmaster:code(50135)]
+description=DRY DEP FLUX: ALICE (3D)
+help=Dry Deposition flux of ALICE (3D)
+ =moles/s
+
+[stashmaster:code(50136)]
+description=WET DEP FLUX: BOB (3D)
+help=Wet Deposition flux of BOB (3D)
+ =moles/s
+
[stashmaster:code(50140)]
description=DMS + OH => SO2 + MeOO + HCHO
help=Chemical reaction flux for DMS + OH => SO2 + MeOO + HCHO
@@ -24136,6 +24151,21 @@
=number of moles of Ox being produced by this reaction per second in the
=whole model.
+[stashmaster:code(52134)]
+description=RXN FLUX: ALICE+OH->BOB+Sec_Org PLEV
+help=Flux through ALICE+OH->BOB+Sec_Org reaction on pressure levels
+ =moles/s
+
+[stashmaster:code(52135)]
+description=DRY DEP FLUX: ALICE ON PRESS LEVELS
+help=Dry Deposition flux of ALICE (3D) on pressure levels
+ =moles/s
+
+[stashmaster:code(52136)]
+description=WET DEP FLUX: BOB ON PRESS LEVELS
+help=Wet Deposition flux of BOB (3D) on pressure levels
+ =moles/s
+
[stashmaster:code(52140)]
description=DMS+OH=>SO2+MeOO+HCHO ON PRESS LEVS
help=Chemical reaction flux for DMS + OH => SO2 + MeOO + HCHO
Index: rose-meta/um-atmos/HEAD/etc/stash/STASHmaster/STASHmaster_A
===================================================================
--- rose-meta/um-atmos/HEAD/etc/stash/STASHmaster/STASHmaster_A (revision 46696)
+++ rose-meta/um-atmos/HEAD/etc/stash/STASHmaster/STASHmaster_A (revision 46718)
@@ -23558,6 +23558,24 @@
4| 1 | 0 | -99 -99 -99 -99 -99 -99 -99 -99 -99 -99 |
5| 0 | 1871 | 0 | 65 | 0 | 0 | 0 | 0 | 0 |
#
+1| 1 | 50 | 134 |RXN FLUX: ALICE+OH->BOB+Sec_Org |
+2| 0 | 0 | 17 | 1 | 2 | 10 | 11 | 0 | 0 | 0 | 0 |
+3| 000000000000000000000000000000 | 00000000000000000001 | 3 |
+4| 1 | 0 | -99 -99 -99 -99 -99 -99 -99 -99 -99 -99 |
+5| 0 | 1871 | 0 | 65 | 0 | 0 | 0 | 0 | 0 |
+#
+1| 1 | 50 | 135 |DRY DEP FLUX: ALICE (3D) |
+2| 0 | 0 | 17 | 1 | 2 | 10 | 11 | 0 | 0 | 0 | 0 |
+3| 000000000000000000000000000000 | 00000000000000000001 | 3 |
+4| 1 | 0 | -99 -99 -99 -99 -99 -99 -99 -99 -99 -99 |
+5| 0 | 1871 | 0 | 65 | 0 | 0 | 0 | 0 | 0 |
+#
+1| 1 | 50 | 136 |WET DEP FLUX: BOB (3D) |
+2| 0 | 0 | 17 | 1 | 2 | 10 | 11 | 0 | 0 | 0 | 0 |
+3| 000000000000000000000000000000 | 00000000000000000001 | 3 |
+4| 1 | 0 | -99 -99 -99 -99 -99 -99 -99 -99 -99 -99 |
+5| 0 | 1871 | 0 | 65 | 0 | 0 | 0 | 0 | 0 |
+#
1| 1 | 50 | 140 |DMS + OH => SO2 + MeOO + HCHO |
2| 0 | 0 | 17 | 1 | 2 | 10 | 11 | 0 | 0 | 0 | 0 |
3| 000000000000000000000000000000 | 00000000000000000001 | 3 |
@@ -26042,6 +26060,24 @@
4| 1 | 0 | -99 -99 -99 -99 -99 -99 -99 -99 -99 -99 |
5| 0 | 1871 | 0 | 8 | 0 | 0 | 0 | 0 | 0 |
#
+1| 1 | 52 | 134 |RXN FLUX: ALICE+OH->BOB+Sec_Org PLEV|
+2| 0 | 0 | 17 | 1 | 3 | 1 | 2 | 0 | 0 | 0 | 1 |
+3| 000000000000000000000000000000 | 00000000000000000001 | 3 |
+4| 1 | 0 | -99 -99 -99 -99 -99 -99 -99 -99 -99 -99 |
+5| 0 | 1871 | 0 | 8 | 0 | 0 | 0 | 0 | 0 |
+#
+1| 1 | 52 | 135 |DRY DEP FLUX: ALICE ON PRESS LEVELS |
+2| 0 | 0 | 17 | 1 | 3 | 1 | 2 | 0 | 0 | 0 | 1 |
+3| 000000000000000000000000000000 | 00000000000000000001 | 3 |
+4| 1 | 0 | -99 -99 -99 -99 -99 -99 -99 -99 -99 -99 |
+5| 0 | 1871 | 0 | 8 | 0 | 0 | 0 | 0 | 0 |
+#
+1| 1 | 52 | 136 |WET DEP FLUX: BOB ON PRESS LEVELS |
+2| 0 | 0 | 17 | 1 | 3 | 1 | 2 | 0 | 0 | 0 | 1 |
+3| 000000000000000000000000000000 | 00000000000000000001 | 3 |
+4| 1 | 0 | -99 -99 -99 -99 -99 -99 -99 -99 -99 -99 |
+5| 0 | 1871 | 0 | 8 | 0 | 0 | 0 | 0 | 0 |
+#
1| 1 | 52 | 140 |DMS+OH=>SO2+MeOO+HCHO ON PRESS LEVS |
2| 0 | 0 | 17 | 1 | 3 | 1 | 2 | 0 | 0 | 0 | 1 |
3| 000000000000000000000000000000 | 00000000000000000001 | 3 |
These differences can be found in the file /home/ukca/Tutorial/vn10.9/worked_solutions/Task9.1/Task9.1_code.patch on PUMA.
If you open the .pa file in Xconv, you should see the following fields:
20 : 96 72 38 1 molfluxd: Stash code = 50134 21 : 96 72 38 1 molfluxd: Stash code = 50135 22 : 96 72 38 1 molfluxd: Stash code = 50136
Sample output from this task can be found at /work/n02/n02/ukca/Tutorial/vn10.9/sample_output/Task9.1/atmosa.pa19810901_00 on ARCHER.
Checklist
- ☐ Make the required changes to branch's STASHmaster_A file, to add the new diagnostics. Make a note of the STASH items chosen, and also add the pressure-level diagnostics.
- ☐ Add help text for your diagnostics in your branch's STASHmaster-meta.conf file.
- ☐ Using a text editor, open the
app/um/rose-app.conffile from yourroses/[SUITE-ID]directory, and add the lineSTASHMASTER=STASHmasterin the[env]block, then save and close the file. - ☐ Using a text editor, open the
rose-suite.conffile from yourroses/[SUITE-ID]directory, and add the following lines to the top of the file, before saving and closing it:
[file:app/um/file/STASHmaster] source=fcm:um.xm_br/dev/[your MOSRS userid]/vnX.Y_your_branch_name/rose-meta/um-atmos/vn10.9/etc/stash/STASHmaster@HEAD
- ☐ Edit your suite using
rose edit -M /path/to/your/branch/working/copy. - ☐ Point the metadata in your suite to um-atmos/vn10.9.
- ☐ Include your branch in your suite at: fcm_make env Sources.
- ☐ In
asad_flux_dat.F90, make a new array of typeasad_flux_defnand populate it with your new diagnostic specification(s), referencing the same item numbers as in your STASHmaster_A file. - ☐ Append this new array at the end of
asad_chemical_fluxes, and incrementn_chemical_fluxesby the number of new diagnostics. - ☐ If required, add new code for new diagnostics into
asad_chem_flux_diags.F90and other UKCA routines as necessary. - ☐ Output your diagnostics in STASH at: um namelist Model Input and Output STASH Requests and Profiles STASH Requests.
- ☐ Run the TidyStashTransform transform macro.
- ☐ Save your suite.
- ☐ In the
roses/[SUITE-ID]directory, runfcm committo commit your changes to the repository. - ☐ Run your suite.
Written by Luke Abraham 2017